Immunoreceptor tyrosine-based activation motif
An immunoreceptor tyrosine-based activation motif (ITAM) is a conserved sequence of four amino acids that is repeated twice in the cytoplasmic tails of non-catalytic tyrosine-phosphorylated receptors, cell-surface proteins found mainly on immune cells.[1] Its major role is being an integral component for the initiation of a variety of signaling pathway and subsequently the activation of immune cells, although different functions have been described, for example an osteoclast maturation.[2][3]
Structure[edit]
The motif contains a tyrosine separated from a leucine or isoleucine by any two other amino acids, giving the signature YxxL/I.[1] Two of these signatures are typically separated by between 6 and 8 amino acids in the cytoplasmic tail of the molecule (YxxL/Ix(6-8)YxxL/I). However, it is worth noting that in various sources, this consensus sequence differs, mainly in the number of amino acids between individual signatures. Apart from ITAMs which have the structure described above, there is also a variety of proteins containing ITAM-like motifs, which have a very similar structure and function (for example in Dectin-1 protein).[4][5][6]
Function[edit]
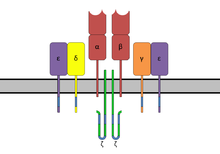
ITAMs are important for signal transduction, mainly in immune cells. They are found in the cytoplasmic tails of non-catalytic tyrosine-phosphorylated receptors[7] such as the CD3 and ζ-chains of the T cell receptor complex, the CD79-alpha and -beta chains of the B cell receptor complex, and certain Fc receptors.[1][7] The tyrosine residues within these motifs become phosphorylated by Src family kinases following interaction of the receptor molecules with their ligands. Phosphorylated ITAMs serve as docking sites for other proteins containing a SH2 domain, usually two domains in tandem, inducing a signaling cascade mediated by Syk family kinases (which are the primary proteins that bind to phosphorylated ITAMs), namely either Syk or ZAP-70, resulting mostly in the activation of given cell. Paradoxically, in some cases, ITAMs and ITAM-like motifs do not have an activating effect, but rather an inhibitory one.[8][9][10] Exact mechanisms of this phenomenon are as of yet not elucidated.
Other non-catalytic tyrosine-phosphorylated receptors carry a conserved inhibitory motif (ITIM) that, when phosphorylated, results in the inhibition of the signaling pathway via recruitment of phosphatases, namely SHP-1, SHP-2 and SHIP1. This serves not only for inhibition and regulation of signalling pathways related to ITAM-based signalling, but also for termination of signalling.[11][12][13]
Genetic variations[edit]
Rare human genetic mutations are catalogued in the human genetic variation databases[14][15][16] which can reportedly result in creation or deletion of ITIM and ITAMs.[17]
Examples[edit]
Examples shown below list both proteins that contain the ITAM themselves and proteins that use ITAM-based signalling with the help of associated proteins which contain the motif.
CD3γ, CD3δ, CD3ε, TYROBP (DAP12), FcαRI, FcγRI, FcγRII, FcγRIII, Dectin-1, CLEC-1, CD28, CD72
References[edit]
- ^ a b c Abbas AK, Lichtman AH (2009), Basic Immunology: Functions and Disorders of the Immune System (3 ed.), Philadelphia, PA: Saunders, ISBN 978-1-4160-4688-2
- ^ Humphrey, Mary Beth; Daws, Michael R.; Spusta, Steve C.; Niemi, Eréne C.; Torchia, James A.; Lanier, Lewis L.; Seaman, William E.; Nakamura, Mary C. (February 2006). "TREM2, a DAP12-associated receptor, regulates osteoclast differentiation and function" (PDF). Journal of Bone and Mineral Research. 21 (2): 237–245. doi:10.1359/JBMR.051016. ISSN 0884-0431. PMID 16418779. S2CID 34957715.
- ^ Paloneva, Juha; Mandelin, Jami; Kiialainen, Anna; Böhling, Tom; Prudlo, Johannes; Hakola, Panu; Haltia, Matti; Konttinen, Yrjö T.; Peltonen, Leena (2003-08-18). "DAP12/TREM2 Deficiency Results in Impaired Osteoclast Differentiation and Osteoporotic Features". Journal of Experimental Medicine. 198 (4): 669–675. doi:10.1084/jem.20030027. ISSN 0022-1007. PMC 2194176. PMID 12925681.
- ^ Rogers, Neil C.; Slack, Emma C.; Edwards, Alexander D.; Nolte, Martijn A.; Schulz, Oliver; Schweighoffer, Edina; Williams, David L.; Gordon, Siamon; Tybulewicz, Victor L.; Brown, Gordon D.; Reis e Sousa, Caetano (April 2005). "Syk-dependent cytokine induction by Dectin-1 reveals a novel pattern recognition pathway for C type lectins". Immunity. 22 (4): 507–517. doi:10.1016/j.immuni.2005.03.004. ISSN 1074-7613. PMID 15845454.
- ^ Underhill, David M.; Rossnagle, Eddie; Lowell, Clifford A.; Simmons, Randi M. (2005-10-01). "Dectin-1 activates Syk tyrosine kinase in a dynamic subset of macrophages for reactive oxygen production". Blood. 106 (7): 2543–2550. doi:10.1182/blood-2005-03-1239. ISSN 0006-4971. PMC 1895265. PMID 15956283.
- ^ Suzuki-Inoue, Katsue; Fuller, Gemma L. J.; García, Angel; Eble, Johannes A.; Pöhlmann, Stefan; Inoue, Osamu; Gartner, T. Kent; Hughan, Sascha C.; Pearce, Andrew C.; Laing, Gavin D.; Theakston, R. David G. (2006-01-15). "A novel Syk-dependent mechanism of platelet activation by the C-type lectin receptor CLEC-2". Blood. 107 (2): 542–549. doi:10.1182/blood-2005-05-1994. ISSN 0006-4971. PMID 16174766. S2CID 168505.
- ^ a b Dushek O, Goyette J, van der Merwe PA (November 2012). "Non-catalytic tyrosine-phosphorylated receptors". Immunological Reviews. 250 (1): 258–76. doi:10.1111/imr.12008. PMID 23046135. S2CID 1549902.
- ^ Pasquier, Benoit; Launay, Pierre; Kanamaru, Yutaka; Moura, Ivan C.; Pfirsch, Séverine; Ruffié, Claude; Hénin, Dominique; Benhamou, Marc; Pretolani, Marina; Blank, Ulrich; Monteiro, Renato C. (January 2005). "Identification of FcalphaRI as an inhibitory receptor that controls inflammation: dual role of FcRgamma ITAM". Immunity. 22 (1): 31–42. doi:10.1016/j.immuni.2004.11.017. ISSN 1074-7613. PMID 15664157.
- ^ O’Neill, Shannon K.; Getahun, Andrew; Gauld, Stephen B.; Merrell, Kevin T.; Tamir, Idan; Smith, Mia J.; Dal Porto, Joseph M.; Li, Quan-Zhen; Cambier, John C. (2011-11-23). "Monophosphorylation of CD79a and b ITAM motifs initiates a SHIP-1 phosphatase-mediated inhibitory signaling cascade required for B cell anergy". Immunity. 35 (5): 746–756. doi:10.1016/j.immuni.2011.10.011. ISSN 1074-7613. PMC 3232011. PMID 22078222.
- ^ Pfirsch-Maisonnas, Séverine; Aloulou, Meryem; Xu, Ting; Claver, Julien; Kanamaru, Yutaka; Tiwari, Meetu; Launay, Pierre; Monteiro, Renato C.; Blank, Ulrich (2011-04-19). "Inhibitory ITAM Signaling Traps Activating Receptors with the Phosphatase SHP-1 to Form Polarized "Inhibisome" Clusters". Science Signaling. 4 (169): ra24. doi:10.1126/scisignal.2001309. ISSN 1945-0877. PMID 21505186. S2CID 206670699.
- ^ Long, Eric O. (August 2008). "Negative signaling by inhibitory receptors: the NK cell paradigm". Immunological Reviews. 224: 70–84. doi:10.1111/j.1600-065X.2008.00660.x. ISSN 1600-065X. PMC 2587243. PMID 18759921.
- ^ Kane, Barry A.; Bryant, Katherine J.; McNeil, H. Patrick; Tedla, Nicodemus T. (2014). "Termination of Immune Activation: An Essential Component of Healthy Host Immune Responses". Journal of Innate Immunity. 6 (6): 727–738. doi:10.1159/000363449. ISSN 1662-811X. PMC 6741560. PMID 25033984.
- ^ Ligeti, E.; Csépányi-Kömi, R.; Hunyady, L. (April 2012). "Physiological mechanisms of signal termination in biological systems". Acta Physiologica. 204 (4): 469–478. doi:10.1111/j.1748-1716.2012.02414.x. ISSN 1748-1716. PMID 22260256. S2CID 13455093.
- ^ Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, et al. (October 2015). "A global reference for human genetic variation". Nature. 526 (7571): 68–74. Bibcode:2015Natur.526...68T. doi:10.1038/nature15393. PMC 4750478. PMID 26432245.
- ^ Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K (January 2001). "dbSNP: the NCBI database of genetic variation". Nucleic Acids Research. 29 (1): 308–11. doi:10.1093/nar/29.1.308. PMC 29783. PMID 11125122.
- ^ Cummings BB, Karczewski KJ, Kosmicki JA, Seaby EG, Watts NA, Singer-Berk M, et al. (May 2020). "Transcript expression-aware annotation improves rare variant interpretation". Nature. 581 (7809): 452–458. Bibcode:2020Natur.581..452C. doi:10.1038/s41586-020-2329-2. PMC 7334198. PMID 32461655.
- ^ Ulaganathan VK (May 2020). "TraPS-VarI: Identifying genetic variants altering phosphotyrosine based signalling motifs". Scientific Reports. 10 (1): 8453. Bibcode:2020NatSR..10.8453U. doi:10.1038/s41598-020-65146-2. PMC 7242328. PMID 32439998.
