Gyrase
| DNA gyrase | ||
|---|---|---|
|
||
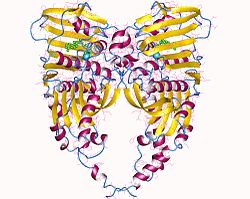
| Gyrase molecule | ||
| Enzyme classification | ||
| EC, category | 5.99.1.3 , topoisomerase | |
| Substrate | DNA | |
A gyrase (. V . AltGr : γῦρος, gyros = circle rounding) is an enzyme , the spatial orientation of closed DNA - molecules changed. It belongs to the topoisomerases type II, which cause a double-strand break in the DNA in prokaryotic cells depending on ATP . The gyrase ensures negative supercoiling - an "unwinding" of the DNA in contrast to the normal structure (B-DNA) with around ten base pairs per turn. This leads to both a gain in space and a partially better readability of the DNA.
Since gyrase = topoisomerase II in this form (with this protein structure) only occurs in bacteria , gyrase inhibitors are also used as antibiotics . The affinity of the gyrase inhibitors for bacterial gyrase is higher than that for human topoisomerases. However, the mechanism of action is not 100% selective and so gyrase inhibitors also have cytostatic properties.
The binding of the ATP to the gyrase is blocked by the antibiotic novobiocin .
A reverse gyrase carries out positive DNA supercoiling , which increases the temperature resistance of the DNA double strand. Their functionality corresponds to that of a type I topoisomerase, coupled with a helicase . Reverse gyrases have been found in bacteria and archaea. Generally they consist of a single polypeptide , with the exception of Methanopyrus kandleri (two parts).
See also
literature
- DB Wigley, GJ Davies, EJ Dodson, A. Maxwell, G. Dodson: Crystal structure of an N-terminal fragment of the DNA gyrase B protein. In: Nature. Volume 351, Number 6328, June 1991, pp. 624-629, doi : 10.1038 / 351624a0 . PMID 1646964 .
- JH Morais Cabral, AP Jackson, CV Smith, N. Shikotra, A. Maxwell, RC Liddington: Crystal structure of the breakage-reunion domain of DNA gyrase. In: Nature. Volume 388, Number 6645, August 1997, pp. 903-906, doi : 10.1038 / 42294 . PMID 9278055 .
- MA Dar, A. Sharma, N. Mondal, SK Dhar: Molecular cloning of apicoplast-targeted Plasmodium falciparum DNA gyrase genes: unique intrinsic ATPase activity and ATP-independent dimerization of PfGyrB subunit. In: Eukaryotic cell. Volume 6, number 3, March 2007, pp. 398-412, doi : 10.1128 / EC.00357-06 . PMID 17220464 . PMC 1828931 (free full text).
- A. Dar, D. Prusty, N. Mondal, SK Dhar: A unique 45-amino-acid region in the top domain of Plasmodium falciparum gyrase B is essential for its activity. In: Eukaryotic cell. Volume 8, Number 11, November 2009, pp. 1759-1769, doi : 10.1128 / EC.00149-09 . PMID 19700639 . PMC 2772398 (free full text).
Web links
- Nomenclature Committee of the International Union of Biochemistry and Molecular Biology (NC-IUBMB): Enzyme Nomenclature. Recommendations. EC 5.99.1.3: DNA topoisomerase (ATP hydrolysing).
- ExPASy: DNA topoisomerase (ATP hydrolysing) .
Individual evidence
- ↑ InterPro: Reverse gyrase
- ↑ Tao-shih Hsieh and Jody L. Plank: Reverse Gyrase Functions as a DNA Renaturase , in: JBC 281 pp. 5640–5647 of January 3, 2006, DOI: 10.1074 / jbc.M513252200