Phosphoglycerate mutase
| Phosphoglycerate mutase | ||
|---|---|---|

|
||
| Phosphoglycerate mutase 1 (B type) dimer (human) according to PDB 1YFK | ||
| Properties of human protein | ||
| Mass / length primary structure | 253/252/254/270 amino acids | |
| Secondary to quaternary structure | Homodimer | |
| Identifier | ||
| Gene names | PGAM1 ; PGAM2 ; PGAM4 ; PGAM5 | |
| Enzyme classification | ||
| EC, category | 5.4.2.1 , isomerase | |
| Response type | Rearrangement | |
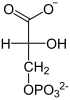
| Substrate | 2-phosphoglycerate | |
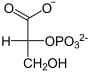
| Products | 3-phosphoglycerate | |
| Occurrence | ||
| Parent taxon | Creature | |
Phosphoglycerate mutases (PGAM) are enzymes that catalyze the rearrangement of the phosphate group in phosphoglycerate from the 2- to 3-position and vice versa . This reaction is a sub-step in glycolysis , the utilization of carbohydrates in the metabolism of all living things, which takes place in every cell . In mammals , additional alleles of the PGAM1 gene called PGAM2, PGAM4 and PGAM5 have formed by copying . PGAM2 is only produced in the muscles. Mutations in the PGAM2 gene lead to a form of muscular dystrophy .
Catalyzed reaction
2-phosphoglycerate is rearranged to 3-phosphoglycerate and vice versa. The catalysis takes place through the interim formation of a phosphohistidine residue in the PGAM. It should be noted that an additional phosphate residue is attached to C2 and thus the intermediate product 2,3-BPG is created.
Only then is the phosphate residue at C3 removed and the mutase receives the phosphoryl group back in order to regenerate the phosphohistidine residue.
The enzyme requires catalytic amounts of 2,3-bisphosphoglycerate in order to keep the histidine residue in the active site in phosphorylated form.
More functions
The PGAM isoforms also have weak activity as bisphosphoglycerate mutase ( EC 5.4.2.4 ) and as bisphosphoglycerate phosphatase ( EC 3.1.3.13 ).
PGAM1 is produced in excess in cancer cells. PGAM1 binds to the core of the hepatitis C virus in vitro . Patients with autoimmune hepatitis produce increased levels of PGAM1 antibodies.
PGAM5 is transported to the outer membrane of mitochondria , where it binds Keap1 and Nrf2 .
Web links
Individual evidence
- ↑ Jeremy M. Berg, John L. Tymoczko, Lubert Stryer, Gregory J. Gatto, Jr.: Stryer Biochemistry . Springer-Verlag, 2014, ISBN 978-3-8274-2988-9 , pp. 467 ( limited preview in Google Book search).
- ↑ F. Lu et al .: Serum proteomic-based analysis for the identification of a potential serological marker for autoimmune hepatitis. Biochem Biophys Res Commun . 367/2/ 2008 : 284-90. PMID 18154727
- ↑ HX Su et al .: Screening cellular proteins binding to the core region of hepatitis C virus RNA genome with digoxin-labeled nucleic acids. Intervirology. 50/4/ 2007 : 303-9. PMID 17622790
- ↑ LJ Huang et al .: Proteomic analysis of secreted proteins of non-small cell lung cancer. Ai Zheng. 25/11/ 2006 : 1361-7. PMID 17094902
- ↑ SC Lo and M. Hannink: PGAM5 tethers a ternary complex Containing Keap1 and Nrf2 to the mitochondria. Exp Cell Res . 314/8/ 2008 : 1789-803. PMID 18387606