Mitochondrion
| Parent |
| Organelle |
| Subordinate |
|
Membrane intermembrane space Matrix |
| Gene Ontology |
|---|
| QuickGO |
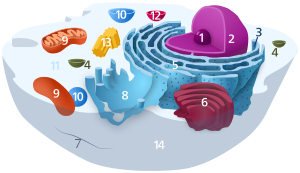
1. Nucleolus (nuclear body)
2. Cell nucleus (nucleus)
3. Ribosomes
4. Vesicle
5. Rough (granular) ER (ergastoplasm)
6. Golgi apparatus
7. Cytoskeleton
8. Smooth (agranular) ER
9 . mitochondria
10. lysosome
11. cytoplasm (with cytosol and cytoskeleton )
12. peroxisomes
13. centrioles
14 cell membrane



• Matrix
• the inner and outer membrane
• diaphragm gap ( intermembrane space ),
• ATP synthase Complexes
• cristae
• mitochondrial ribosome (Mitoribosomen)
• granules
• many existing circular mitochondrial DNA (mtDNA)

The term mitochondrion or mitochondrion (in ancient Greek μίτος mitos 'thread' and χονδρίον chondrion 'granule'; outdated chondriosome ) is a cell organelle that is surrounded by a double membrane and contains its own genetic material, the mitochondrial DNA . Mitochondria occur as spherical or tubular structures in the cells of almost all eukaryotes , but not in prokaryotes .
Mitochondria regenerate the high- energy molecule adenosine triphosphate via the respiratory chain . In addition to this oxidative phosphorylation , they fulfill other essential tasks for the cell, for example they are involved in the formation of iron-sulfur clusters .
General
A particularly large number of mitochondria are located in cells with high energy consumption; These include muscle cells , nerve cells , sensory cells and egg cells . In cardiac muscle cells , the volume fraction of mitochondria reaches 36%. They have a diameter of about 0.5–1.5 µm and have a wide variety of shapes, from spheres to complex networks. Mitochondria reproduce by growth and budding , the number of mitochondria is adapted to the energy requirements of the cell. Eukaryotic cells that lose their mitochondria can no longer regenerate them. There are also eukaryotes without mitochondria, e.g. B. some protozoa . The number of mitochondria per cell is typically on the order of 1000 to 2000 with a volume fraction of 25%, but these values can vary greatly depending on the cell type and organism. The number in the mature human sperm cell is around four to five mitochondria located in the middle (“neck”), whereas in the mature egg cell it is several hundred thousand.
Mitochondria are usually only inherited from the mother via the plasma of the egg cell, which prompted research into maternal kin lines ( matrilines ). Diseases that are caused by mutations in mitochondrial genes are usually only passed on from the mother.
In the meantime it has been found that some male mitochondria are also imported into the plasma of the fertilized egg cell ( zygote ) through the sperm . However, these “male” mitochondria are usually eliminated very quickly. However, there are a few cases where medical professionals have shown that the child's mitochondria were from the paternal line.
So far, about 50 diseases ( mitochondrial diseases ) are known that can be caused by mitochondrial dysfunction.
The German pathologist and histologist Richard Altmann discovered the mitochondrion, 1886. The term "powerhouse of the cell" (often used popular science English powerhouse of the cell ) for the mitochondrion was coined by Philip Siekevitz 1957th
construction
The envelope of the mitochondria consists of an outer and an inner membrane , which are made up of phospholipid bilayers and proteins . Both membranes differ in their properties. Four different compartments exist through these membranes : the outer membrane, the intermembrane space (the space between the two membranes), the inner membrane / the cristae and the matrix (the space within the inner membrane).
Mitochondria are able to connect (by fusion) and to divide again (by fission); in yeast cells, about two mitochondrial fusions or missions occur per minute. It is therefore not possible to precisely determine the current number of mitochondria in a cell.
However, mitochondria do not just occur as separate, bean-shaped organelles. Instead, they can also form tubular, i.e. tubular, and sometimes branched networks. Here form and function are closely related. They are subject to various influences and dynamic regulation. Such microtubular extensions, which are larger than the T-tubules in skeletal muscles and which sometimes connect mitochondria with one another over greater distances, enable an intermitochondrial exchange of electrolytes and a stabilization of the membrane potential . They are mainly observed in mitochondrial myopathies , but can also occur much less frequently in normal muscle cells . Possibly they are used for the survival of the mitochondria under stress, as in stress-induced mitophagy .
Outer membrane
The outer membrane encloses the entire mitochondrion and contains channels made of protein complexes, which enable the exchange of molecules and ions between the mitochondrion and the cytosol . Large molecules cannot pass through the membrane.
The mitochondrial outer membrane, which encloses the entire organelle and is not folded, has a weight ratio of phospholipid to protein of 1: 1 and is therefore similar to the eukaryotic plasma membrane. It contains numerous integral proteins, the porins . Porins form channels that allow molecules with a mass of up to 5000 Daltons to freely diffuse through the membrane. Larger proteins can enter the mitochondria when a signal sequence binds at its N-terminus to a large protein subunit of the translocase , which then actively moves them across the membrane. If cracks appear in the outer membrane, proteins can escape from the intermembrane space into the cytosol, which can lead to cell death. The mitochondrial outer membrane may be that of the endoplasmic reticulum unite (ER), and then forms a structure which MAM ( English mitochondria-associated ER-membrane is called). This is important for signal exchange between the ER and the mitochondrion and plays a role in lipid transfer .
Intermembrane space
The intermembrane space (membrane space) is the space between the outer membrane and the inner membrane. Since the outer membrane is freely permeable to small molecules, the concentration of small molecules such as ions and sugars in the intermembrane space is identical to that in the cytosol . However, large proteins require a specific signal sequence in order to be transported through the membrane, so that the composition of the proteins differs between the intermembrane space and the cytosol. One protein that is held in this way in the intermembrane space is cytochrome c .
Inner membrane
The proteins of the inner membrane can be divided into five groups according to their function:
- Those who perform the redox reactions of oxidative phosphorylation
- ATP synthase , which produce ATP in the matrix
- Special transport proteins that regulate the entry and exit of metabolites from the matrix
- The protein import machinery
- Proteins used to fuse and split mitochondria
The inner membrane contains more than 151 different polypeptides and has a very high protein-to-phospholipid weight ratio (more than 3: 1, which equates to about 15 phospholipids for every 1 protein). Around 1/5 of the total proteins in a mitochondrion are located in the inner membrane. In addition, the inner membrane rich in an unusual phospholipid, the cardiolipin . This was originally discovered in cow hearts in 1942 and is usually characteristic of mitochondrial and bacterial plasma membranes. Cardiolipin contains four fatty acids instead of the usual two, is responsible for the ion impermeability of the membrane and is otherwise only found in prokaryotes . In contrast to the outer membrane, the inner membrane does not contain any porins and is impermeable to all molecules. Almost all ions and molecules therefore require special membrane transporters when entering or leaving the matrix. Proteins are transported into the matrix via the translocase of the inner membrane (TIM) or via Oxa1. In addition, there is a membrane potential between the intermembrane space and the matrix, which is formed by the enzymes in the respiratory chain.
The inner membrane encloses the matrix, the internal fluid of the mitochondrion. It corresponds to the cytosol of bacteria and contains the mitochondrial DNA (mtDNA), the enzymes of the citric acid cycle and its own mitochondrial ribosomes ( mitoribosomes ).
Structure of the inner membrane
The cristae type (from Latin crista "comb") has numerous indentations, the cristae, on the inner membrane. This significantly increases the surface area of the inner membrane where chemical reactions can take place and increases its ability to produce ATP. The inner membrane contains large protein complexes from the respiratory chain , which are responsible for generating energy. The other type of mitochondria is called the tubule type and is found, for example, in steroid-producing cells; his inner membrane forms tubes.
In the case of tubular invaginations with pearl-like round bulges, one speaks of the sacculi type. In the mitochondria of a liver cell, the area of the inner membrane is about five times larger than that of the outer membrane. This ratio is variable, and mitochondria from cells with an increased need for ATP, such as muscle cells, contain even more cristae. The cristae are covered on the inside with small round bodies with a diameter of 8.5 nm, which are known as F1 particles, elementary particles or ATP synthase particles. This is where the formation of ATP takes place in the course of cell respiration . The indentations or folds of the inner membrane are not accidental and can impair their chemiosmotic function.
Mitoplast
Mitoplast is a term for the mitochondrion without an outer membrane: a mitochondrion whose outer membrane has been removed (for examination purposes) so that only an intact inner membrane and its contents (matrix) are left.
matrix
The space enclosed by the inner membrane is called the matrix. It contains 2/3 of all proteins in a mitochondrion. The matrix is important in the production of ATP, which takes place with the help of ATP synthase. The matrix contains a highly concentrated mixture of hundreds of enzymes as well as the special mitochondrial ribosomes (mitoribosomes), tRNA and several copies of the mitochondrial genome (mitogenome). In addition, the concentration of intermediates (intermediate products) of the citric acid cycle of beta oxidation is very high. The main tasks of the enzymes include the oxidation of pyruvate and fatty acids and the citric acid cycle.
Since mitochondria have their own genetic material, they can produce RNA and proteins themselves ( transcription and translation ). It was shown that 16,569 base pairs encode a total of 37 genes in the mitochondrial DNA sequence (mitogenome) , of which 22 are tRNA, 2 rRNA and 13 are peptide genes. The 13 mitochondrial peptides are integrated into the inner mitochondrial membrane, along with proteins that are encoded in the genome of the host cell and are formed by it and then introduced.
The mitochondrial ribosomes (mitoribosomes) differ from the eukaryotic ribosomes in the cytosol. The differences to bacterial ribosomes are comparatively small, but they do exist. Mitochondrial ribosomes in humans are about 55S Svedberg units, compared to 70S in bacteria and 80S in cytosolic ribosomes. Although there are mitochondrial 70S ribosomes, but not in mammals ( mammals ) including humans. The intermembrane space between the two membranes contains enzymes that can phosphorylate nucleotides while consuming ATP .
Mitochondria-associated ER membrane (MAM)
The mitochondria-associated ER membrane (MAM, English mitochondria-associated ER membrane ) is another structural element, its crucial role in cellular physiology and homeostasis is increasingly recognized. While it was once suspected that it was just a permanent impurity from technical difficulties in fractionation, it has now been identified as a membranous structure at the interface between mitochondria and the ER. A physical coupling between these two organelles had previously been investigated using electron microscopy and more recently using fluorescence microscopy . Such investigations led to the assumption that on the MAM, which encloses up to 20% of the outer mitochondrial membrane, ER and mitochondrion are only separated by a 10–25 nm gap and both are held together by a protein complex.
Purified MAM from subcellular fractionations has shown that, in addition to Ca 2+ ion channels, it is also enriched with enzymes that are involved in the phospholipid exchange. These indications of a special role of MAM in the regulation of fat storage and signal transmission have been confirmed, with important implications for mitochondrial-associated cellular phenomena, as discussed below. MAM not only provided an insight into the underlying mechanistic principles of physiological processes such as intrinsic apoptosis and the spread of calcium signals, but it also refined our view of the mitochondria. Although they are often seen as static and isolated "power plants of the cell", the development of MAM underscores the extent to which mitochondria have been integrated into cell physiology, with close physical and functional coupling to the endomembrane system.
Phospholipid transfer
The MAM is enriched with enzymes that are involved in lipid biosynthesis, such as. B. Phosphatidylserine synthase on the ER surface and Phosphatidylserine decarboxylase on the mitochondrial surface. Since mitochondria, as dynamic organelles, undergo constant splitting and fusion, they must be supplied with phospholipids in a constant and adequately regulated manner for membrane integrity. However, mitochondria are not only a target for phospholipids, but also play a role in the exchange between organelles of (intermediate) products of the phospholipid biosynthetic pathways, the ceramide and cholesterol metabolism and glycosphingolipid anabolism.
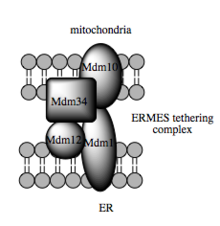
The transport capacity of these substances depends on the MAM, which has been shown to facilitate the transfer of lipid intermediates between organelles. In contrast to the standard vesicle mechanism of lipid transfer, there are indications that the spatial proximity of the ER to the mitochondrial membrane in MAM enables flipping between opposing lipid bilayers. Despite this unusual and apparently energetically unfavorable mechanism, such transport does not require ATP. Instead, a function of a multi-protein complex has been shown, the ERMES ( English ER-mitochondria encounter structure ) is known, even if it remains unclear whether this complex directly gives the lipid transfer or else is required to hold the membranes in close enough proximity to each other to lower the energy barrier for direct lipid flipping.
MAM could also be part of the secretory mechanism in addition to its role in intracellular lipid exchange. In particular, MAM appears to be an intermediate station between the rough ER and the Golgi apparatus, in the metabolic pathway that leads to Very Low Density Lipoproteins (VLDL). The MAM thus serves as a crucial metabolic and transshipment point in lipid metabolism.
Calcium signaling
The important role of the ER in calcium signaling was recognized long before that of the mitochondrion; partly because the low affinity of the Ca 2+ channels in the outer membrane completely contradicted the supposed responsibility of the organelle for intracellular Ca 2+ currents. This apparent contradiction is resolved by the presence of the MAM: the close spatial connection between the two organelles results in Ca 2+ microdomains at points of contact, which facilitate efficient Ca 2+ transfer from the ER to the mitochondria. The transmission takes place in response to so-called “Ca 2+ puffs”, generated by spontaneous aggregation and activation of IP3R , a recognized ER membrane Ca 2+ channel.
perspective
The MAM is of crucial importance for the exchange of information and metabolic products in the cell, which enables the physiology of ER and mitochondria to be networked. There is not only a structural, but also a functional coupling between the two, which is crucial for the entire cellular physiology and homeostasis. The MAM thus enables a view of the mitochondria that deviates from the traditional view of these organelles as a static, isolated unit. Instead, the integration of the mitochondria into different cellular processes is emphasized as the result of an endosymbiotic process.
function
- Important breakdown pathways: Citric acid cycle , for this purpose pyruvate is introduced from the cytosol into the mitochondrial matrix. By the pyruvate dehydrogenase pyruvate is then to acetyl-CoA decarboxylation. Another source of acetyl-CoA is the breakdown of fatty acids ( β-oxidation ), which takes place in the mitochondria in animal cells, but only in the glyoxysomes and peroxisomes in plant cells. For this purpose, acyl-CoA from the cytosol is channeled through the inner mitochondrial membrane by binding to carnitine and converted to acetyl-CoA. In the citric acid cycle (also known as the Krebs cycle or tricarboxylic acid cycle), the majority of the reduction equivalents (NADH + H + , FADH 2 ) are obtained from acetyl-CoA , which are then converted into ATP within the respiratory chain.
- Respiratory chain : With the help of electron transport processes and the enrichment of protons, an electrochemical gradient is built up between the intermembrane space and the mitochondrial matrix, which is used to produce ATP using the ATP synthase (see chemiosmotic coupling ). The electrons and protons required to build up the gradient are obtained from the nutrients (e.g. glucose ) absorbed by the organism through oxidative degradation . First of all, glycolysis occurs in the cytoplasm .
- Apoptosis (programmed cell death)
- Calcium storage: mitochondria intervene in the calcium homeostasis of the cell through the ability to absorb calcium ions and later release them again .
- Synthesis of iron-sulfur clusters , which are required by many enzymes in the respiratory chain , among other things . This function is now considered to be the essential function of the mitochondria, i. H. as the reason almost all eukaryotic cells rely on mitochondria for survival.
- Individual steps from the urea cycle also take place in mitochondria.
origin
According to the endosymbiotic theory , it is assumed that the mitochondria emerged from a symbiosis of aerobic bacteria (from the group of α-proteobacteria , genus Rickettsia ) with the ancestors of today's eukaryotes . An alternative proposal is the uptake of an optional anaerobic bacterium ( symbiont ) by a methanogenic archaeon ( host ). Indications of an endosymbiosis, however designed, are the possession of one's own genetic information (mtDNA, mitogenome or - less often - chondrioma), one's own protein synthesis - with its own ribosomes (mitoribosomes) and its own tRNAs - and the presence of an inner membrane that is clearly different from the The structure of the outer membrane is different and that serves the synthesis of ATP from ADP . However, the mitochondria are so specialized that they are not viable on their own. They are relatively closely related to other, less common organelles , the hydrogenosomes . However, these usually - not always - lack their own DNA, as do the related mitosomes .
Together with the hydrogenosomes and mitosomes, mitochondria are therefore classified as " mitochondrial-related organelles" ( English mitochondrion-related organelles , MROs). These also include the anaerobic and DNA- free organelles of Henneguya salminicola (alias H. zschokkei , Myxozoa )
Almost without exception, all eukaryotes have MROs, i.e. mitochondria or organelles of one of these related types. An exception is Monocercomonoides ( Excavata ). It is believed that these unicellular organisms acquired a cytosolic system through horizontal gene transfer in order to provide the iron-sulfur clusters required for protein synthesis. After that, their mitochondrial organelles were superfluous in all their functions and were lost.
Genome
The mitochondria (almost without exception) have their own genome ( chondrioma , also mitogenome), which is often copied several times and is located in the mitochondrial matrix. The genome is usually present in a single circular, double-stranded DNA (mtDNA) (see also plasmid ), which can, however, be present in several (about two to ten) copies. The mtDNA has an independent duplication cycle. Mitochondria are called semiautonomous; their genome itself only encodes a small part of the proteins required by the mitochondrion . In animals, the mitogenome typically consists of 37 genes 16 kb (base pairs) in length . In humans, the 37 mitochondrial genes control the synthesis of 13 of the approximately 80 protein subunits of the respiratory chain, the remaining 800–1000 different mitochondrial proteins are encoded in the nuclear genome.
The genes of mtDNA that do not code for proteins code for the rRNA and for all required tRNAs .
Changes in the mitochondrial genome are used in research to elucidate lineages of species , as well as different ethnic groups of humans, for example by the Genographic project .
Multiplication
Mitochondria are not newly formed, but arise through growth and sprouting . The majority of mitochondrial proteins are synthesized in the cytosol and then transported into the mitochondria. The transport of these proteins into the mitochondria via the outer membrane by the TOM complex ( English translocase of outer mitochondrial membrane ) and across the inner membrane by the TIM complex (engl. Translocase of inner mitochondrial membrane ) and includes the function of chaperones , especially Hsp70 .
The reproduction of the mitochondria is homologous and highly conserved across all types of eukaryotes .
The extent and timing of mitochondria multiplication depend on requirements. During cell division , they are distributed from the mother cell to the daughter cells.
Used mitochondria are broken down with the help of the endoplasmic reticulum, the Golgi apparatus and the lysosomes .
Mitochondria in complex chloroplasts
Interestingly, there are complex chloroplasts (which, according to the endosymbiotic theory, arise from a secondary endosymbiosis of a eukaryote with an algae cell - for example a red alga ) with their own mitochondria. Some dinoflagellates such as Kryptoperidinium and Durinskia (both Peridiniaceae , also English dinotoms ) have a chloroplast derived from diatoms ( Heterokontophyta ). These chloroplasts are surrounded by up to five membranes, depending on whether the entire diatom endosymbiont is seen as the chloroplast or only the red alga it contains is counted as the chloroplast. The diatom endosymbiont has been reduced relatively little - it still retains its original mitochondria and has endoplasmic reticulum, eukaryotic ribosomes , a cell nucleus and of course the complex (secondary) chloroplasts derived from red algae - practically a complete cell - all inside the host.
The ocelloids of the unicellular dinoflagellates in the family Warnowiaceae (Warnowiiden) are also complex organelles composed of plastids and mitochondria.
Possible relationships with aging
The mitochondria, as the power plants of the cell, can have leaks in the respiratory chain; ie in the case of respiratory oxidation, electrons can be released. These can form reactive oxygen species , which can cause oxidative stress , which can lead to a high mutation rate in mitochondrial DNA (mtDNA). These hypothetical links between aging and oxidative stress are not new and were proposed over 50 years ago. They are questioned in more recent works.
Oxidative stress can lead to mitochondrial DNA mutations, which can cause enzymatic abnormalities, which in turn can lead to further oxidative stress. A vicious circle is possible. However, recent measurements have shown that the rate of accumulation of mutations in mitochondrial DNA is 1 mutation per 7884 years (i.e. 10 −7 to 10 −9 per base per year, considering the youngest common ancestors of humans and monkeys), and is therefore comparable on the mutation rates of autosomal DNA (10 −8 bases per generation).
Various changes can occur during the aging of the mitochondria. Tissues from elderly patients show a decrease in the enzymatic activity of the proteins in the respiratory chain . However, mutations in mitochondrial DNA can only be found in 0.2% of the very old cells. It is believed that large deletions in the mitochondrial genome lead to high levels of oxidative stress and neuronal death, which triggers Parkinson's disease .
However, there is much debate as to whether mitochondrial changes are causes of aging or just characteristics of aging. A representative study on mice showed a shortened lifespan, but no increase in reactive oxygen species despite increasing mitochondrial DNA mutations. It should be noted, however, that aging wild-type mice do not accumulate as many mutations in mitochondrial DNA as previously thought (and these mutations have less of an effect than expected), which raises doubts about the importance of mutations in mitochondrial DNA for natural aging leaves. Thus, the exact connections between mitochondria, oxidative stress and aging remain unexplained.
See also
- Mitochondrial disease
- Mitochondrial Eve
- Hydrogenosome
- Mitosome
- Plastid , especially chloroplasts and complex chloroplasts
- Ocelloid
literature
- Bruce Alberts et al. a .: Molecular Biology of the Cell. 4th edition. 2002, ISBN 0-8153-3218-1 ( ncbi.nlm.nih.gov ).
- MW Gray, G. Burger, BF Lang: The origin and early evolution of mitochondria . In: Genome Biology . tape 2 , no. 6 , January 2001, p. 1018 , PMID 11423013 ( genomebiology.com [PDF]).
- Gottfried Schatz : Mitochondria. Beyond oxidative phosphorylation . In: Biochimica et Biophysica Acta . tape 1271 , no. 1 , May 24, 1995, p. 123-126 , PMID 7599197 .
- IE Scheffler: A century of mitochondrial research: achievements and perspectives . In: Mitochondrion . tape 1 , no. 1 , June 1, 2001, ISSN 1567-7249 , p. 3-31 , PMID 16120266 ( mitoresearch.org [PDF]).
Web links
- Esther Neumann: Power plants of our cells (Top Life Aktuell 10/2005)
- zytologie-online.net: Mitochondrion graphics
- MitoP2 ( Memento from July 14, 2006 in the Internet Archive ) - English-language database for mitochondrial genes, proteins and diseases
- MITOMAP - A human mitochondrial genome database - English language site on the mitochondrial genome
- Uni Mainz: Electron microscopic images of mitochondria
- Wilfried Probst: Early Evolution and Symbiosis , European University Flensburg, Institute for Biology and Science and its Didactics: § LECA and Mitochondria, accessed on April 19, 2019
Individual evidence
- ^ Wilhelm Gemoll : Greek-German school and hand dictionary. Munich / Vienna 1965.
- ↑ Emil Heitz : The structural relationships between plant and animal chondriosomes . In: Journal for Nature Research. 12b / 1957, pp. 576-578, doi: 10.1515 / znb-1957-8-916
- ↑ J. Schrader, M. Kelm: The heart. In: Rainer Klinke, Hans-Christian Pape, Stefan Silbernagl (eds.): Physiology. 3. Edition. Georg Thieme Verlag, Stuttgart 2005, p. 147.
- ↑ Esther Neumann: Power plants of our cells. In: Top Life Aktuell. 10/2005.
- ↑ Jan Koolman: Pocket Atlas of Biochemistry. 3. Edition. Georg Thieme Verlag, Stuttgart 2003, p. 210.
- ^ Science online, compact dictionary of biology: Spermium. Spectrum Academic Publishing House, 2007.
- ^ Rolf Knippers: Molecular Genetics. 9th edition. Thieme Verlag, Konstanz 2006, p. 455.
- ↑ Medical research: British create embryo with three parents. on: stern.de , February 6, 2008. See also Mitochondrial DNA .
- ↑ M. Zeviani, S. di Donato: Mitochondrial disorders. In: Brain. 127 (2004), pp. 2153-2172; PMID 15358637 . (PDF; 789 kB)
- ↑ P. Siekevitz: Power House of the cell . In: Scientific American . 197, No. 1, 1957, pp. 131-140. doi : 10.1038 / scientificamerican0757-131 .
- ↑ a b c d e f g Bruce Alberts, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, Peter Walter: Molecular Biology of the Cell . Garland Publishing, New York 1994, ISBN 0-8153-3218-1 .
- ^ Donald Voet, Judith G. Voet, Charlotte W. Pratt: Fundamentals of Biochemistry . 2nd Edition. John Wiley and Sons, 2006, ISBN 0-471-21495-7 , pp. 547 .
- ↑ Stefan Jakobs: Mitochondria - Dynamic power plants of the cell. (PDF; 4.9 MB); Max Planck Institute for Biophysical Chemistry, Göttingen. In: MPIbpc News. 10 (2004), issue 12, pp. 1-4.
- ↑ SM Rafelski: Mitochondrial network morphology: building an integrative, geometrical view. In: BMC biology. Volume 11, June 2013, p. 71, doi : 10.1186 / 1741-7007-11-71 , PMID 23800141 , PMC 3691739 (free full text) (review).
- ↑ Silverio Perrotta, Domenico Roberti, Debora Bencivenga, Paola Corsetto, Katie A. O'Brien, Martina Caiazza, Emanuela Stampone, Leanne Allison, Roland A. Fleck, Saverio Scianguetta, Immacolata Tartaglione, Peter A. Robbins, Maddalena Casale, James A. West, Clara Franzini-Armstrong, Julian L. Griffin, Angela M. Rizzo, Antonio A. Sinisi, Andrew J. Murray, Adriana Borriello, Federico Formenti, Fulvio Della Ragione: Effects of Germline VHL Deficiency on Growth, Metabolism, and Mitochondria . New England Journal of Medicine 2020, Volume 382, Issue 9, February 27, 2020, Pages 835-844, DOI: 10.1056 / NEJMoa1907362
- ↑ a b J. M. Herrmann, W. Neupert: Protein transport into mitochondria . In: Curr Opin Microbiol . tape 3 , no. 2 , April 2000, p. 210-214 , doi : 10.1016 / S1369-5274 (00) 00077-1 , PMID 10744987 .
- ↑ a b J. E. Chipuk, L. Bouchier-Hayes, DR Green: Mitochondrial outer membrane permeabilization during apoptosis: the innocent bystander scenario . In: Cell Death and Differentiation . tape 13 , no. 8 , 2006, p. 1396-1402 , doi : 10.1038 / sj.cdd.4401963 , PMID 16710362 .
- ^ T. Hayashi, R. Rizzuto, G. Hajnoczky, TP Su: MAM: more than just a housekeeper . In: Trends Cell Biol . tape 19 , no. 2 , February 2009, p. 81-88 , doi : 10.1016 / j.tcb.2008.12.002 , PMID 19144519 , PMC 2750097 (free full text).
- ↑ JB McMillin, W. Dowhan: Cardiolipin and apoptosis . In: Biochim. Biophys. Acta . tape 1585 , no. 2–3 , December 2002, pp. 97-107 , doi : 10.1016 / S1388-1981 (02) 00329-3 , PMID 12531542 .
- ↑ a b Katharina Munk, Konstanze Abröll, Thomas Kurth, Thomas Langer, Regina Nethe-Jaenchen, Harald Schlatter, Beate Schultze, Klaus Wolf: Biochemistry - Cell Biology . 1st edition. Thieme, 2008, ISBN 3-13-144831-8 , p. 433 .
- ^ CA Mannella: Structure and dynamics of the mitochondrial inner membrane cristae . In: Biochimica et Biophysica Acta (BBA) - Mol Cell Res . tape 1763 , no. 5-6 , 2006, pp. 542-548 , doi : 10.1016 / j.bbamcr.2006.04.006 , PMID 16730811 .
- ↑ Farlex Free dictionary ; ScienceDirect
- ↑ S. Anderson, AT Bankier, BG Barrell, MHL de-Bruijn, AR Coulson et al. : Sequence and organization of the human mitochondrial genome . In: Nature . tape 290 , no. 5806 , 1981, pp. 427-465 , doi : 10.1038 / 290457a0 , PMID 7219534 .
- ↑ Rassow et al. : Biochemistry. Duale series, 2nd edition, Thieme, p. 446.
- ↑ TW O'Brien: Properties of human mitochondrial ribosomes . In: IUBMB Life . tape 55 , no. 9 , September 2003, p. 505-513 , doi : 10.1080 / 15216540310001626610 , PMID 14658756 .
- ↑ a b c d e f g R. Rizzuto et al. : Ca2 + transfer from the ER to mitochondria: when, how and why . In: Biochim Biophys Acta . tape 1787 , no. 11 , 2009, p. 1342-1351 , doi : 10.1016 / j.bbabio.2009.03.015 , PMID 19341702 , PMC 2730423 (free full text).
- ↑ a b T. Hayashi et al. : MAM: more than just a housekeeper . In: Trends Cell Biol . tape 19 , no. 2 , 2009, p. 81-88 , doi : 10.1016 / j.tcb.2008.12.002 , PMID 19144519 , PMC 2750097 (free full text).
- ↑ a b O. M. de Brito et al. : An intimate liaison: spatial organization of the endoplasmic reticulum – mitochondria relationship . In: EMBO J. Band 29 , no. 16 , 2010, p. 2715–2723 , doi : 10.1038 / emboj.2010.177 , PMID 20717141 , PMC 2924651 (free full text).
- ↑ a b J. E. Vance et al. : Intracellular trafficking of phospholipids: import of phosphatidylserine into mitochondria . In: Anticancer Research . tape 16 , 3B, 1996, pp. 1333-1339 , PMID 8694499 .
- ↑ a b c M. Lebiedzinska et al. : Interactions between the endoplasmic reticulum, mitochondria, plasma membrane and other subcellular organelles . In: Int J Biochem Cell Biol . tape 41 , no. 10 , 2009, p. 1805-1816 , doi : 10.1016 / j.biocel.2009.02.017 , PMID 19703651 .
- ↑ G. Twig et al. : Fission and selective fusion govern mitochondrial segregation and elimination by autophagy . In: EMBO J . tape 27 , no. 2 , 2008, p. 433-446 , doi : 10.1038 / sj.emboj.7601963 , PMID 18200046 , PMC 2234339 (free full text).
- ↑ a b c d e C. Osman et al. : Making heads or tails of phospholipids in mitochondria . In: J Cell Biol . tape 192 , no. 1 , 2011, p. 7–16 , doi : 10.1083 / jcb.201006159 , PMID 21220505 , PMC 3019561 (free full text).
- ↑ B. Kornmann et al. : An ER-Mitochondria Tethering Complex Revealed by a Synthetic Biology Screen . In: Science . tape 325 , no. 24 , 2009, p. 477-481 , doi : 10.1126 / science.1175088 , PMID 19556461 , PMC 2933203 (free full text).
- ↑ AE Rusinol et al. : A Unique Mitochondria-associated Membrane Fraction from Rat Liver Has a High Capacity for Lipid Synthesis and Contains Pre-Golgi Secretory Proteins Including Nascent Lipoprotein . In: J Biol Chem . tape 269 , no. 44 , 1994, pp. 27494-27502 , PMID 7961664 .
- ↑ Prof. Weiler, Lutz Nover: General and Molecular Botany. Thieme, Stuttgart a. a. 2008, ISBN 978-3-13-147661-6 , p. 80.
- ↑ R. Lill et al. a .: The essential ro <nowike /> le of mitochondria in the biogenesis of cellular iron-sulfur proteins. In: Biological Chemistry . Volume 380 (1999), pp. 1157-1166. PMID 10595578
- ↑ W. Martin, MJ Russell: On the Origin of Cells: a Hypothesis for the Evolutionary Transitions from Abiotic Geochemistry to Chemoautotropic Prokaryotes, and from Prokaryotes to Nucleated Cells. In: Philos Trans R Soc Lond B Biol Sci. 358 (1429), Jan 23, 2003, pp. 59-83; Discussion pp. 83-85.
- ↑ B. Boxma et al. a .: An anaerobic mitochondrion that produces hydrogen. In: Nature. Volume 434 (2005), No. 7029, pp. 74-79. PMID 15744302 , doi: 10.1038 / nature03343
- ↑ Jan Osterkamp: First animal without breathing and mitochondria , on: Spektrum.de from February 25, 2020
- ↑ Tel Aviv University researchers discover unique non-oxygen breathing animal , on: EurekAlert! from February 25, 2020
- ↑ See also: H. nuesslini ; Peacock Wrasse§ Threats ( H. tunisiensis )
- ↑ Anna Karnkowska, Vojtěch Vacek, Zuzana Zubáčová, Sebastian C. Treitli, Romana Petrželková, Laura Eme, Lukáš Novák, Vojtěch Žárský, Lael D. Barlow, Emily K. Herman, Petr Soukal, Miluše Hroudová, Pavel Doležal., Courtney W. Stairs , Andrew J. Roger, Marek Eliáš, Joel B. Dacks, Čestmír Vlček, Vladimír Hampl: A Eukaryote without a Mitochondrial Organelle . In: Current Biology . 26, No. 10, 2016, ISSN 0960-9822 , pp. 1274-1284. doi : 10.1016 / j.cub.2016.03.053 . PMID 27185558 .
- ↑ Davis, Josh L .: Scientists Shocked to Discover Eukaryote With NO Mitochondria . IFL Science. May 13, 2016. Retrieved April 9, 2019.
- ↑ RJ Wiesner, JC Rüegg, I. Morano: Counting target molecules by exponential polymerase chain reaction: copy number of mitochondrial DNA in rat tissues . In: Biochemical and Biophysical Research Communications . 183, No. 2, March 1992, pp. 553-559. doi : 10.1016 / 0006-291X (92) 90517-O . PMID 1550563 .
- ↑ J. M. Herrmann: Protein transport machines in mitochondria. In: Naturwissenschaftliche Rundschau . 58 (2005), pp. 525-530.
- ↑ SciTechDaily: Breakthrough in Understanding Evolution - Mitochondrial Division Conserved Across Species , December 30, 2019, source: Tokyo University of Science
- ↑ a b c P. J. Keeling: The endosymbiotic origin, diversification and fate of plastids . In: Philosophical Transactions of the Royal Society B: Biological Sciences . 365, No. 1541, 2010, pp. 729-48. doi : 10.1098 / rstb.2009.0103 . PMID 20124341 . PMC 2817223 (free full text).
- ↑ a b Patrick J. Keeling: Diversity and evolutionary history of plastids and their hosts . In: American Journal of Botany . 91, No. 10, 2004, pp. 1481-93. doi : 10.3732 / ajb.91.10.1481 . PMID 21652304 .
- ↑ Jacques Joyard, Maryse A. Block, Roland Douce: Molecular aspects of plastid envelope biochemistry . In: Eur. J. Biochem. . 199, No. 3, 1991, pp. 489-509. doi : 10.1111 / j.1432-1033.1991.tb16148.x . PMID 1868841 .
- ↑ Chloroplast . In: Encyclopedia of Science . Retrieved March 20, 2019.
- ↑ Eberhard Schnepf, Malte Elbrächter: Dinophyte chloroplasts and phylogeny - A review . In: Grana . 38, No. 2-3, 1999, pp. 81-97. doi : 10.1080 / 00173139908559217 .
- ↑ C. Richter, J. Park, BN Ames: Normal oxidative damage to mitochondrial and nuclear DNA is extensive . In: PNAS . tape 85 , no. September 17 , 1988, pp. 6465-6467 , doi : 10.1073 / pnas.85.17.6465 , PMID 3413108 , PMC 281993 (free full text).
- ^ D. Harman: Aging: a theory based on free radical and radiation chemistry . In: J. Gerontol . tape 11 , no. 3 , 1956, pp. 298-300 , PMID 13332224 .
- ↑ D. Gems, L. Partridge: Genetics of Longevity in Model Organisms: Debates and Paradigm Shifts . In: Annu Rev Physiol. tape 75 , 2012, p. 25.1-25.24 , doi : 10.1146 / annurev-physiol-030212-183712 , PMID 23190075 .
- ↑ Soares et al. : Correcting for Purifying Selection: An Improved Human Mitochondrial Molecular Clock. In: Am J Hum Genet. 84 (6), June 12, 2009, pp. 740-759, doi: 10.1016 / j.ajhg.2009.05.001 , PMC 2694979 (free full text).
- ↑ Michael W. Nachman, Susan L. Crowell: Estimate of the Mutation Rate per Nucleotide in Humans. In: Genetics. Vol. 156, September 2000, pp. 297-304.
- ↑ D. Boffoli, SC Scacco, R. Vergari, G. Solarino, G. Santacroce, S. Papa: Decline with age of the respiratory chain activity in human skeletal muscle . In: Biochim. Biophys. Acta . tape 1226 , no. 1 , 1994, p. 73-82 , doi : 10.1016 / 0925-4439 (94) 90061-2 , PMID 8155742 .
- ↑ Aubrey de Gray: Mitochondrial Mutations in Mammalian Aging: An Over-Hasty About-Turn? ( Memento of the original from October 21, 2007 in the Internet Archive ) Info: The archive link was automatically inserted and not yet checked. Please check the original and archive link according to the instructions and then remove this notice. In: Rejuvenation Res. 7 (3), Fall 2004, pp. 171-174. doi: 10.1089 / rej.2004.7.171 . PMID 15588517 .
- ↑ A. Bender, KJ Krishnan, CM Morris, GA Taylor, AK Reeve, RH Perry, E. Jaros, JS Hersheson, J. Betts, T. Klopstock, RW Taylor, DM Turnbull: High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson's disease . In: Nat Gen . tape 38 , no. 5 , 2006, p. 515-517 , doi : 10.1038 / ng1769 , PMID 16604074 .
- Jump up ↑ A. Trifunovic, A. Hansson, A. Wredenberg, AT Rovio, E. Dufour, I. Khvorostov, JN Spelbrink, R. Wibom, HT Jacobs, NG Larsson: Somatic mtDNA mutations cause aging phenotypes without affecting reactive oxygen species production . In: PNAS . tape 102 , no. 50 , 2005, pp. 17993-17998 , doi : 10.1073 / pnas.0508886102 , PMID 16332961 , PMC 1312403 (free full text).
- ↑ Marc Vermulst et al. : Mitochondrial point mutations do not limit the natural lifespan of mice . In: Nature Genetics . tape 39 , no. 4 , 2007, p. 540-543 , doi : 10.1038 / ng1988 , PMID 16332961 .