Metamonad: Difference between revisions
wrong url |
Animalculum (talk | contribs) mNo edit summary |
||
| Line 32: | Line 32: | ||
==Characteristics== |
==Characteristics== |
||
A number of parabasalids and oxymonads are found in [[termite]] [[guts]], and play an important role in breaking down the [[cellulose]] found in [[wood]]. |
A number of parabasalids and oxymonads are found in [[termite]] [[guts]], and play an important role in breaking down the [[cellulose]] found in [[wood]]. Some other metamonads are [[Parasitism|parasites]]. |
||
These flagellates are unusual in lacking [[mitochondrion|mitochondria]]. |
These flagellates are unusual in lacking [[mitochondrion|mitochondria]]. Originally they were considered among the most primitive [[eukaryote]]s, diverging from the others before mitochondria appeared. However, they are now known to have lost mitochondria secondarily, and retain both organelles and nuclear genes derived from them. Mitochondrial relics include [[hydrogenosome]]s, which produce [[hydrogen]], and small structures called [[mitosome]]s. |
||
It now appears the Metamonada are, together with ''[[Malawimonas]]'', sister clades of the [[Unikont|Podiata]].<ref>{{Cite journal|last1=Cavalier-Smith|first1=Thomas|last2=Chao|first2=Ema E.|last3=Lewis|first3=Rhodri|date=2016-06-01|title=187-gene phylogeny of protozoan phylum Amoebozoa reveals a new class (Cutosea) of deep-branching, ultrastructurally unique, enveloped marine Lobosa and clarifies amoeba evolution|journal=Molecular Phylogenetics and Evolution|volume=99|pages=275–296|doi=10.1016/j.ympev.2016.03.023|pmid=27001604|doi-access=free}}</ref> |
It now appears the Metamonada are, together with ''[[Malawimonas]]'', sister clades of the [[Unikont|Podiata]].<ref>{{Cite journal|last1=Cavalier-Smith|first1=Thomas|last2=Chao|first2=Ema E.|last3=Lewis|first3=Rhodri|date=2016-06-01|title=187-gene phylogeny of protozoan phylum Amoebozoa reveals a new class (Cutosea) of deep-branching, ultrastructurally unique, enveloped marine Lobosa and clarifies amoeba evolution|journal=Molecular Phylogenetics and Evolution|volume=99|pages=275–296|doi=10.1016/j.ympev.2016.03.023|pmid=27001604|doi-access=free}}</ref> |
||
All of these groups are united by having [[flagellum|flagella]] or basal bodies in characteristic groups of four, which are often associated with the [[cell nucleus|nucleus]], forming a structure called a |
All of these groups are united by having [[flagellum|flagella]] or basal bodies in characteristic groups of four, which are often associated with the [[cell nucleus|nucleus]], forming a structure called a karyomastigont. In addition, the genera ''[[Carpediemonas]]'' and ''[[Trimastix]]'' are now known to be close relatives of the retortamonad-diplomonad line and the oxymonads, respectively. Both are free-living and amitochondriate. |
||
== Classification == |
== Classification == |
||
The metamonads were thought to make up part of the [[Excavata]], a paraphyletic eukaryotic supergroup including flagellates with feeding grooves and their close relatives. |
The metamonads were thought to make up part of the [[Excavata]], a paraphyletic eukaryotic supergroup including flagellates with feeding grooves and their close relatives. Their relationships are uncertain,<ref name="pmid14657102">{{cite journal |author=Cavalier-Smith T |title=The excavate protozoan phyla Metamonada Grassé emend. (Anaeromonadea, Parabasalia, Carpediemonas, Eopharyngia) and Loukozoa emend. (Jakobea, Malawimonas): their evolutionary affinities and new higher taxa |journal=Int. J. Syst. Evol. Microbiol. |volume=53 |issue=Pt 6 |pages=1741–58 |date=November 2003 |pmid=14657102 |doi= 10.1099/ijs.0.02548-0|doi-access=free }}</ref> and they do not always appear together on molecular trees. It is possible that the metamonads as defined here do not form a [[monophyletic]] subgroup. |
||
The following higher level treatment from 2013 is based on works of Cavalier-Smith<ref>{{cite journal |author=Cavalier-Smith T |title=Early evolution of eukaryote feeding modes, cell structural diversity, and classification of the protozoan phyla Loukozoa, Sulcozoa, and Choanozoa |journal= Eur. J. Protistol. |volume=49 |issue=2 |pages=115–178 |date= 2013 |pmid=23085100 |doi=10.1016/j.ejop.2012.06.001 }}</ref> with amendments within Fornicata from Yubukia, Simpson & Leander.<ref>{{cite journal |author1=Yubukia |author2=Simpson |author3=Leander |title=Comprehensive Ultrastructure of Kipferlia bialata Provides Evidence for Character Evolution within the Fornicata (Excavata) |journal=Protist |volume= 164|issue= 3|pages= 423–439|date= 2013 |pmid= 23517666|doi=10.1016/j.protis.2013.02.002 }}</ref> |
The following higher level treatment from 2013 is based on works of [[Thomas Cavalier-Smith|Cavalier-Smith]]<ref>{{cite journal |author=Cavalier-Smith T |title=Early evolution of eukaryote feeding modes, cell structural diversity, and classification of the protozoan phyla Loukozoa, Sulcozoa, and Choanozoa |journal= Eur. J. Protistol. |volume=49 |issue=2 |pages=115–178 |date= 2013 |pmid=23085100 |doi=10.1016/j.ejop.2012.06.001 }}</ref> with amendments within [[Fornicata]] from Yubukia, Simpson & Leander.<ref>{{cite journal |author1=Yubukia |author2=Simpson |author3=Leander |title=Comprehensive Ultrastructure of Kipferlia bialata Provides Evidence for Character Evolution within the Fornicata (Excavata) |journal=Protist |volume= 164|issue= 3|pages= 423–439|date= 2013 |pmid= 23517666|doi=10.1016/j.protis.2013.02.002 }}</ref> |
||
Metamonada were once again proposed to be basal eukaryotes in 2018.<ref>{{Cite journal|last1=Krishnan|first1=Arunkumar|last2=Burroughs|first2=A. Max |last3=Iyer |first3=Lakshminarayan|last4=Aravind|first4=L.|date=2018-07-04|title=The unexpected provenance of components in eukaryotic nucleotide-excision-repair and kinetoplast DNA-dynamics from bacterial mobile elements|url=https://www.biorxiv.org/content/early/2018/07/04/361121|journal=bioRxiv |pages=361121 |doi=10.1101/361121 |doi-access=free}}</ref> |
Metamonada were once again proposed to be basal eukaryotes in 2018.<ref>{{Cite journal|last1=Krishnan|first1=Arunkumar|last2=Burroughs|first2=A. Max |last3=Iyer |first3=Lakshminarayan|last4=Aravind|first4=L.|date=2018-07-04|title=The unexpected provenance of components in eukaryotic nucleotide-excision-repair and kinetoplast DNA-dynamics from bacterial mobile elements|url=https://www.biorxiv.org/content/early/2018/07/04/361121|journal=bioRxiv |pages=361121 |doi=10.1101/361121 |doi-access=free}}</ref> |
||
Latest revision as of 20:32, 14 April 2024
| Metamonad | |
|---|---|
| |
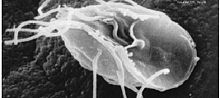
| Giardia lamblia, a parasitic diplomonad | |
| Scientific classification | |
| Domain: | Eukaryota |
| (unranked): | Excavata |
| Phylum: | Metamonada Grassé 1952 emend. Cavalier-Smith 2003 |
| Classes & orders | |
| Synonyms[2] | |
| |
The metamonads are a large group of flagellate amitochondriate microscopic eukaryotes. Their composition is not entirely settled, but they include the retortamonads, diplomonads, and possibly the parabasalids and oxymonads as well. These four groups are all anaerobic (many being aerotolerant anaerobes), occurring mostly as symbiotes or parasites of animals, as is the case with Giardia lamblia which causes diarrhea in mammals.[2]
Characteristics[edit]
A number of parabasalids and oxymonads are found in termite guts, and play an important role in breaking down the cellulose found in wood. Some other metamonads are parasites.
These flagellates are unusual in lacking mitochondria. Originally they were considered among the most primitive eukaryotes, diverging from the others before mitochondria appeared. However, they are now known to have lost mitochondria secondarily, and retain both organelles and nuclear genes derived from them. Mitochondrial relics include hydrogenosomes, which produce hydrogen, and small structures called mitosomes.
It now appears the Metamonada are, together with Malawimonas, sister clades of the Podiata.[3]
All of these groups are united by having flagella or basal bodies in characteristic groups of four, which are often associated with the nucleus, forming a structure called a karyomastigont. In addition, the genera Carpediemonas and Trimastix are now known to be close relatives of the retortamonad-diplomonad line and the oxymonads, respectively. Both are free-living and amitochondriate.
Classification[edit]
The metamonads were thought to make up part of the Excavata, a paraphyletic eukaryotic supergroup including flagellates with feeding grooves and their close relatives. Their relationships are uncertain,[4] and they do not always appear together on molecular trees. It is possible that the metamonads as defined here do not form a monophyletic subgroup.
The following higher level treatment from 2013 is based on works of Cavalier-Smith[5] with amendments within Fornicata from Yubukia, Simpson & Leander.[6]
Metamonada were once again proposed to be basal eukaryotes in 2018.[7]
- Phylum Metamonada (Grassé 1952) Cavalier-Smith 1987 emend. Cavalier-Smith 2003
- Family Anaeramoebidae Táborský, Pánek & Čepička 2017
- Subphylum Anaeromonada Cavalier-Smith 1997 emend. 2003
- Class Anaeromonadea Cavalier-Smith 1997 emend. 1999
- Family Paratrimastigidae Zhang et al. 2015[8]
- Order Trimastigida Cavalier-Smith 2003
- Family Trimastigidae Saville Kent 1880
- Order Oxymonadida Grassé 1952 emend. Cavalier-Smith 2003
- Family Polymastigidae Bütschli 1884
- Family Saccinobaculidae Brugerolle & Lee 2002 ex Cavalier-Smith 2013
- Family Pyrsonymphidae Grassé 1892
- Family Oxymonadidae Kirby 1928
- Class Anaeromonadea Cavalier-Smith 1997 emend. 1999
- Subphylum Trichozoa Cavalier-Smith 1996 emend. Cavalier-Smith 2003 stat. n. 2013
- Superclass Fornicata Simpson 2003 stat. n. Cavalier-Smith 2013
- Family Kipferliidae Cavalier-Smith 2013
- Class Carpediemonadea Cavalier-Smith 2013 s.s.
- Order Carpediemonadida Cavalier-Smith 2003 emend. 2013 s.s.
- Family Carpediemonadidae Cavalier-Smith 2003
- Order Carpediemonadida Cavalier-Smith 2003 emend. 2013 s.s.
- Class Eopharyngea Cavalier-Smith 1993 stat. n. Cavalier-Smith 2003
- Order Dysnectida Cavalier-Smith 2013
- Family Dysnectidae Cavalier-Smith 2013
- Order Retortamonadida Grassé 1952 emend. Cavalier-Smith 2013
- Family Caviomonadidae Cavalier-Smith 2013
- Family Chilomastigidae Cavalier-Smith 2013
- Family Retortamonadidae Wenrich 1932
- Order Diplomonadida Wenyon 1926 emend. Brugerolle et al. 1975
- Family Giardiidae Kulda & Nohy´nkova´ 1978
- Family Octomitidae Cavalier-Smith 1996
- Family Spironucleidae Cavalier-Smith 1996
- Family Hexamitidae Kent 1880 emend. Brugerolle et al. 1975
- Order Dysnectida Cavalier-Smith 2013
- Superclass Parabasalia Honigberg 1973 stat. n. Cavalier-Smith 2003
- Class Trichonymphea Cavalier-Smith 2003
- Order Lophomonadida Light 1927
- Family Lophomonadidae Saville Kent 1880
- Order Trichonymphida Poche 1913
- Family †Burmanymphidae Poinar 2009
- Family Retractinymphidae Radek & Brune 2023[9]
- Family Spirotrichosomidae Hollande & Caruette-Valentin 1971
- Family Staurojoeninidae Grassé 1917
- Family Trichonymphidae Saville Kent 1880
- Family Hoplonymphidae Light 1926
- Family Teratonymphidae Koidzumi 1921 [Eucomonymphidae]
- Order Lophomonadida Light 1927
- Class Trichomonadea Kirby 1947 stat. n. Cavalier-Smith 2003
- Order Pimpavickida Céza & Čepička 2022[10]
- Family Pimpavickidae Céza & Čepička 2022
- Order Trichomonadida Kirby 1947
- Family Lacusteriidae Céza & Čepička 2022[10]
- Family Trichomonadidae Chalmers & Pekkoloa 1918 sensu Hampl et al. 2006
- Order Honigbergiellida Čepička et al. 2010[11][10]
- Family Honigbergiellidae Čepička, Hampl & Kulda 2010
- Family Hexamastigidae Čepička, Hampl & Kulda 2010
- Family Tricercomitidae Čepička, Hampl & Kulda 2010
- Order Hypotrichomonadida Čepička et al. 2010
- Family Hypotrichomonadidae (Honigberg 1963) Čepička, Hampl & Kulda 2010
- Order Spirotrichonymphida Grassé 1952
- Family Spirotrichonymphidae Grassé 1917
- Order Tritrichomonadida Čepička et al. 2010
- Family Dientamoebidae Grassé 1953
- Family Monocercomonadidae Kirby 1944
- Family Simplicimonadidae Čepička et al. 2010
- Family Tritrichomonadidae Honigberg 1963
- Order Cristamonadida Brugerolle & Patterson 2001 emend. Cavalier-Smith 2013
- Family Calonymphidae Grassé 1911
- Family Devescovinidae Doflein 1911
- Order Pimpavickida Céza & Čepička 2022[10]
- Class Trichonymphea Cavalier-Smith 2003
- Superclass Fornicata Simpson 2003 stat. n. Cavalier-Smith 2013
References[edit]
- ^ Stairs, Courtney W.; Táborský, Petr; Salomaki, Eric D.; Kolisko, Martin; Pánek, Tomáš; Eme, Laura; Hradilová, Miluše; Vlček, Čestmír; Jerlström-Hultqvist, Jon; Roger, Andrew J.; Čepička, Ivan (2021-12-20). "Anaeramoebae are a divergent lineage of eukaryotes that shed light on the transition from anaerobic mitochondria to hydrogenosomes". Current Biology. 31 (24): 5605–5612.e5. doi:10.1016/j.cub.2021.10.010. ISSN 0960-9822. PMID 34710348. S2CID 240054026.
- ^ a b Al Jewari, Caesar; Baldauf, Sandra L. (2023-04-28). "An excavate root for the eukaryote tree of life". Science Advances. 9 (17): eade4973. Bibcode:2023SciA....9E4973A. doi:10.1126/sciadv.ade4973. ISSN 2375-2548. PMC 10146883. PMID 37115919.
- ^ Cavalier-Smith, Thomas; Chao, Ema E.; Lewis, Rhodri (2016-06-01). "187-gene phylogeny of protozoan phylum Amoebozoa reveals a new class (Cutosea) of deep-branching, ultrastructurally unique, enveloped marine Lobosa and clarifies amoeba evolution". Molecular Phylogenetics and Evolution. 99: 275–296. doi:10.1016/j.ympev.2016.03.023. PMID 27001604.
- ^ Cavalier-Smith T (November 2003). "The excavate protozoan phyla Metamonada Grassé emend. (Anaeromonadea, Parabasalia, Carpediemonas, Eopharyngia) and Loukozoa emend. (Jakobea, Malawimonas): their evolutionary affinities and new higher taxa". Int. J. Syst. Evol. Microbiol. 53 (Pt 6): 1741–58. doi:10.1099/ijs.0.02548-0. PMID 14657102.
- ^ Cavalier-Smith T (2013). "Early evolution of eukaryote feeding modes, cell structural diversity, and classification of the protozoan phyla Loukozoa, Sulcozoa, and Choanozoa". Eur. J. Protistol. 49 (2): 115–178. doi:10.1016/j.ejop.2012.06.001. PMID 23085100.
- ^ Yubukia; Simpson; Leander (2013). "Comprehensive Ultrastructure of Kipferlia bialata Provides Evidence for Character Evolution within the Fornicata (Excavata)". Protist. 164 (3): 423–439. doi:10.1016/j.protis.2013.02.002. PMID 23517666.
- ^ Krishnan, Arunkumar; Burroughs, A. Max; Iyer, Lakshminarayan; Aravind, L. (2018-07-04). "The unexpected provenance of components in eukaryotic nucleotide-excision-repair and kinetoplast DNA-dynamics from bacterial mobile elements". bioRxiv: 361121. doi:10.1101/361121.
- ^ Zhang, Qianqian; Táborský, Petr; Silberman, Jeffrey D.; Pánek, Tomáš; Čepička, Ivan; Simpson, Alastair G.B. (2015). "Marine Isolates of Trimastix marina Form a Plesiomorphic Deep-branching Lineage within Preaxostyla, Separate from Other Known Trimastigids (Paratrimastix n. gen.)". Protist. 166 (4): 468–491. doi:10.1016/j.protis.2015.07.003. PMID 26312987.
- ^ Radek, Renate; Platt, Katja; Öztas, Deniz; Šobotník, Jan; Sillam-Dussès, David; Hanus, Robert; Brune, Andreas (26 January 2023). "New insights into the coevolutionary history of termites and their gut flagellates: Description of Retractinympha glossotermitis gen. nov. sp. nov. (Retractinymphidae fam. nov.)". Frontiers in Ecology and Evolution. 11. doi:10.3389/fevo.2023.1111484.
- ^ a b c Céza, Vít; Kotyk, Michael; Kubánková, Aneta; Yubuki, Naoji; Šťáhlavský, František; Silberman, Jeffrey D.; Čepička, Ivan (August 2022). "Free-living Trichomonads are Unexpectedly Diverse". Protist. 173 (4): 125883. doi:10.1016/j.protis.2022.125883. PMID 35660751. S2CID 248586911.
- ^ Cepicka, Ivan; Hampl, Vladimír; Kulda, Jaroslav (July 2010). "Critical Taxonomic Revision of Parabasalids with Description of one New Genus and three New Species". Protist. 161 (3): 400–433. doi:10.1016/j.protis.2009.11.005. PMID 20093080.
