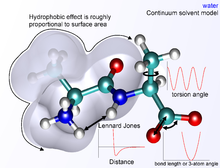
Protein design
The Protein Design , synonym protein engineering or rational protein design , refers to the targeted adjustment of properties of proteins by site-directed mutagenesis of the DNA . Alongside randomly directed evolution, it is one of the two strategies of protein engineering .
principle
The goals of protein design are changes in
- Expression levels
- Binding properties (eg. As substrate affinity , substrate specificity , affinity for other binding partners)
- catalytic properties (e.g. metabolism , substrate saturation )
- toxic properties
- immunological properties (e.g. repetitive epitopes , MHC binding , consensus sequences of different strains, masking by glycosylation )
- the localization in a cell compartment
- in the case of inclusion bodies, the solubility
- Increase in the biological half-life by reducing proteolysis and increasing denaturation and thermal stability
Process and Effects
The targeted modification of recombinant proteins can lead to loss or gain of functionality. In addition to targeted changes to protein, DNA sections outside of the protein-coding sequence are usually also changed in the course of a vector design to increase gene expression . The gene expression can be increased by choosing a promoter , enhancer and terminator suitable for the respective species . Furthermore, a Shine-Dalgarno sequence (in bacteria) or a Kozak sequence (in eukaryotes) can improve the recognition of the mRNA on the ribosome , and a polyadenylation signal at the 3 'end and avoidance of AUUUA sequences can prevent premature degradation the mRNA are reduced.
Point mutations
By codon optimization expression rate can be increased by only those 20 amino acid codons are used which are most highly expressed in the particular type (see codon usage ). The frequent use of suboptimal codons, on the other hand, is a method of attenuating live viral vaccines . In addition to the codons, other RNA sequences can also affect the amount of protein formed and are included in codon optimization. Post-translationally modifiable amino acids, such as those found in glycosylation, phosphorylation , methylation , acetylation , sulfation , myristylation , palmitoylation , farnesylation , GPI anchor and geranylgeranylation sites, can be introduced into the protein through targeted point mutations in the DNA or removed.
By changing a catalytic center , a substrate binding site or a binding site for other molecules that is necessary for activation (e.g. in the case of cofactors , temporary protein-protein interactions or in protein complexes ), competitive inhibitors can be generated.
The biological half-life of a protein may be extended in the Peptidaseschnittstellen , PEST sequences and specific N -terminal amino acids from the N-end rule be changed.
Point mutations can affect the secondary, tertiary and quaternary structure, such as, among other things also on the change in the primary structure of disulfide bridges -ausbildende cysteines . α-Helices can be modified by rotationally flexible ( glycine ), helix-forming (e.g. alanine ) and helix-breaking amino acids ( proline ). Unusual amino acids can be introduced through the use of an extended genetic code .
Insertions and deletions
New protein domains and the associated functions can be added by inserting DNA sequences (made up of multiples of three nucleotides) into a gene in conformance with the reading grid . The resulting hybrid proteins are known as fusion proteins .
Occasionally, for purification and detection, short DNA sequences conforming to the reading frame are inserted after the start codon or before the stop codon of the gene, which are referred to as protein tags .
Further customary insertions are flexible connections ( linker , between two protein domains of a fusion protein ), as well as inteins or protease recognition sequences which enable part of the protein to be cleaved in vitro or in vivo .
Transient insertions can be created by inserting inteins or by using the Cre-lox system .
Properties can be removed by reading grid-compliant deletions of multiples of three nucleotides. Other properties of the protein can occasionally come to the fore, such as B. in the removal of regulatory domains. The localization in a cell compartment can be changed by adding or removing signal sequences . By adding or removing a transmembrane domain , soluble proteins and membrane proteins can be converted into one another. When coding sequences for cell-penetrating peptides are inserted , the cell entry of a protein can be increased.
Multimerization
By modifying viral capsid proteins or by multimerizing proteins, larger protein particles can be produced.
stabilization
Various small proteins (mostly less than 200 amino acids) are used as a framework for stabilization, e.g. B. Affibodies , Affimers , ankyrin repeat proteins ( DARPins ), Repebodies , Anticalins , Fibronectins and Kunitz protein domains . Cyclopeptides are closed ring-shaped peptides which, due to cyclization, have no ends and have a longer biological half-life. Proteins can be stabilized by cross-linking , e.g. B. in an immobilization . Β-peptides generated by peptide synthesis have an elongated peptide backbone.
coupling
By crosslinking agents, for. B. two proteins are coupled to each other in vitro .
mark
Various signal molecules can be attached to a protein in the course of marking.
Examples
In the early 21st century, the development of protein design accelerated through the use of molecular modeling on the computer . Examples of this development include stereoselective catalysis , ion detection , and antiviral properties.
In 2003, computer-aided methods were used to create a new, artificial protein fold (Top7), which also resulted in sensors for unnatural molecules. The specificity for xylose reductase cofactors of Candida boidinii was also changed from NADPH to NADH .
However, it is likely that not all protein structures can be obtained by protein design, since some configurations and conformations cannot develop for steric reasons. There are also software-based limits to the possibilities for change.
software
- IPRO alters the proteins to increase their affinity for a substrate or cofactor. This is achieved through several random changes in the protein backbone in the area of specific positions to identify the lowest energy combinations of the rotamers and to determine the configuration with the lowest energy in the event of a specific change. The iterative approach allows IPRO the additive calculation of several mutations to optimize substrate specificity or cofactor binding.
- EGAD: A Genetic Algorithm for Protein Design . A free software package for protein design and predicting the effects of mutations on protein folding and affinity. EGAD also uses several structures in parallel when designing binding sites or fixed conformations. Movable ligands with or without rotating bonds can also be calculated. EGAD can also be used with multiple processors.
- RosettaDesign . A software package that is free for academic use. RosettaDesign is available through a web server.
- Sharpen is an open source library for protein design and structure prediction. SHARPEN offers various combinatorial optimization methods (e.g. Monte Carlo, Simulated Annealing, FASTER) and evaluates the proteins using the '' Rosetta all-atom force field "or the '' molecular mechanics force field '' (OPLSaa). SHARPEN also includes also the possibility of calculating with several processors.
- WHAT IF software . A software for modeling, protein design, validation and visualization of proteins.
- CheShift is software for the validation of protein structures.
- Abalone is a software for modeling and visualization.
- ProtDes is a software for protein design based on the `` CHARMM molecular mechanics package ''.
literature
- Friedrich Lottspeich , Haralabos Zorbas: Bioanalytics . Spectrum Academic Publishing House, Heidelberg 1998. ISBN 978-3-8274-0041-3 .
- Hubert Rehm , Thomas Letzel: The Experimenter: Protein Biochemistry / Proteomics . 6th edition, Spektrum Akademischer Verlag, Heidelberg 2009. ISBN 978-3-8274-2312-2 .
- E. Buxbaum: Fundamentals of Protein Structure and Function , Springer, New York 2007. ISBN 978-0-387-26352-6 .
- P. Kaumaya: Protein Engineering , Intech, 2012. ISBN 978-953-510-037-9 . Online version (English)
- KP Murphy: Methods In Molecular Biology, Vol. 168: Protein Structure, Stability, And Folding , Humana, New Jersey 2001. ISBN 978-0-89603-682-6 .
- T. Vo-Dinh: Methods in Molecular Biology, Vol. 300: Protein Nanotechnology, Protocols, Instrumentation, and Applications , Humana, New Jersey 2004. ISBN 978-1-58829-310-7 .
- R. Guerois, M. de la Paz: Methods in Molecular Biology, Vol. 340: Protein Design , Humana, New Jersey 2006. ISBN 1-59745-116-9 .
- K. Arndt, K. Muller: Methods in Molecular Biology, Vol. 352: Protein Engineering Protocols , Humana, New Jersey 2007. ISBN 978-1-58829-072-4 .
- Kevin M. Ulmer: Protein engineering. In: Science (1983) 219: 666-71 (4585). PMID 6572017 .
Individual evidence
- ↑ E. Kotsopoulou, VN Kim, AJ Kingsman, SM Kingsman, KA Mitrophanous: A Rev-independent human immunodeficiency virus type 1 (HIV-1) -based vector that exploits a codon-optimized HIV-1 gag-pol gene. In: J Virol. (2000), Vol. 74 (10), pp. 4839-52. PMID 10775623 ; PMC 112007 (free full text).
- ↑ S. Mueller, JR Coleman, E. Wimmer: Putting synthesis into biology: a viral view of genetic engineering through de novo gene and genome synthesis. In: Chemistry & biology. Volume 16, number 3, March 2009, pp. 337-347, doi : 10.1016 / j.chembiol.2009.03.002 , PMID 19318214 , PMC 2728443 (free full text).
- ^ S. Fath, AP Bauer, M. Liss, A. Spriestersbach, B. Maertens, P. Hahn, C. Ludwig, F. Schäfer, M. Graf, R. Wagner: Multiparameter RNA and codon optimization: a standardized tool to assess and enhance autologous mammalian gene expression. In: PLOS ONE . Volume 6, number 3, March 2011, p. E17596, doi : 10.1371 / journal.pone.0017596 , PMID 21408612 , PMC 3048298 (free full text).
- ^ R. Martin: Methods in Molecular Biology, Vol. 77: Protein Synthesis , Humana, New Jersey 1998. ISBN 978-0-89603-397-9 .
- ↑ Yen-Ting Lai, Eamonn Reading, Greg L. Hura, Kuang-Lei Tsai, Arthur Laganowsky, Francisco J. Asturias, John A. Tainer, Carol V. Robinson, Todd O. Yeates: Structure of a designed protein cage that self -assembles into a highly porous cube. In: Nature Chemistry. 2014, S., doi : 10.1038 / nchem.2107 .
- ↑ A. Skerra: Alternative non-antibody scaffolds for molecular recognition. In: Current Opinion in Biotechnology. Volume 18, number 4, August 2007, pp. 295-304, doi : 10.1016 / j.copbio.2007.04.010 . PMID 17643280 .
- ↑ M. Gebauer, A. Skerra: Engineered protein scaffolds as next-generation antibody therapeutics. In: Current opinion in chemical biology. Volume 13, Number 3, June 2009, pp. 245-255, doi : 10.1016 / j.cbpa.2009.04.627 . PMID 19501012 .
- ↑ Alan Saghatelian, Yohei Yokobayashi, Kathy Soltani, M. Reza Ghadiri: A chiroselective peptide replicator. In: Nature. 409, pp. 797-801, doi : 10.1038 / 35057238 .
- ↑ T. Nagai: Circularly permuted green fluorescent proteins engineered to sense Ca2 +. In: Proceedings of the National Academy of Sciences . 98, pp. 3197-3202, doi : 10.1073 / pnas.051636098 .
- ^ MJ Root: Protein Design of an HIV-1 Entry Inhibitor. In: Science. 291, pp. 884-888, doi : 10.1126 / science.1057453 .
- ^ B. Kuhlman: Design of a Novel Globular Protein Fold with Atomic-Level Accuracy. In: Science. 302, 2003, pp. 1364-1368, doi : 10.1126 / science.1089427 .
- ↑ Loren L. Looger, Mary A. Dwyer, James J. Smith, Homme W. Hellinga: Computational design of receptor and sensor proteins with novel functions. In: Nature. 423, 2003, pp. 185-190, doi : 10.1038 / nature01556 .
- ^ A b George A. Khoury, Hossein Fazelinia, Jonathan W. Chin, Robert J. Pantazes, Patrick C. Cirino, Costas D. Maranas: Computational design of xylose reductase for altered cofactor specificity. In: Protein Science. 18, 2009, pp. 2125-2138, doi : 10.1002 / pro.227 .
- ↑ User's Manual for EGAD! a Genetic Algorithm for Protein Design! . Archived from the original on May 2, 2009. Retrieved October 17, 2013.
- ^ Y. Liu, B. Kuhlman: RosettaDesign server for protein design. In: Nucleic Acids Research . 34, 2006, pp. W235-W238, doi : 10.1093 / nar / gkl163 .
- ↑ Gautam Dantas, Brian Kuhlman, David Callender, Michelle Wong, David Baker: A Large Scale Test of Computational Protein Design: Folding and Stability of Nine Completely Redesigned Globular Proteins. In: Journal of Molecular Biology . 332, 2003, pp. 449-460, doi : 10.1016 / S0022-2836 (03) 00888-X .
- ↑ Gautam Dantas, Colin Corrent, Steve L. Reichow, James J. Havranek, Ziad M. Eletr, Nancy G. Isern, Brian Kuhlman, Gabriele Varani, Ethan A. Merritt, David Baker: High-resolution Structural and Thermodynamic Analysis of Extreme Stabilization of Human Procarboxypeptidase by Computational Protein Design. In: Journal of Molecular Biology. 366, 2007, pp. 1209-1221, doi : 10.1016 / j.jmb.2006.11.080 .
- ↑ Rosetta Design . Retrieved August 20, 2012.
- ↑ SHARPEN . Retrieved August 20, 2012.
- ^ Johan Desmet, Jan Spriet, Ignace Lasters: Fast and accurate side-chain topology and energy refinement (FASTER) as a new method for protein structure optimization. In: Proteins: Structure, Function, and Genetics. 48, 2002, pp. 31-43, doi : 10.1002 / prot.10131 .
- ↑ Abalone . Retrieved August 20, 2012.
- ↑ PROTDES . Retrieved August 20, 2012.