Cell disruption
As cell disruption (or homogenization ) in the are biochemistry method referred to, in which cells to be destroyed in order for their content ( homogenate ) - organelle , proteins , DNA , mRNA or other biomolecules - proceed.
Mechanical digestion process
Physical digestion processes usually lead to an increase in temperature, which is why the temperature must be controlled and, if necessary, the digestion material must be cooled in an ice bath or the digestion must be carried out intermittently. Animal cells and plant cells have to be freed of unwanted tissue parts before they are disrupted by roughly chopping them up with scissors, scalpel or meat grinder. In addition, the tissue to be disrupted is shredded by multicellular cells in a blender with rotating knives (or an Ultra-Turrax for small volumes) before homogenization .
- The Potter-Elvehjem method or Dounce method is used when the cell organelles are to be preserved. A piston that is tightly encased in a stationary vessel moves. The cells are destroyed by shear forces .
- In the case of microorganisms , but also all other cells, small volumes can be ground in a mortar with sand or corundum or in a glass bead mill.
- With ultrasound , cavitation forces constantly push and shear the cells against one another. Chromosomal DNA breaks down in the middle because of its mechanical natural oscillation . In all of these processes, a lot of heat is generated when treating larger volumes.
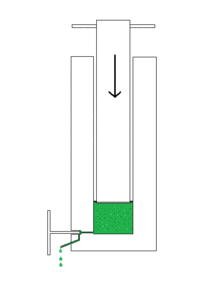
- For continuous operation or volumes> 1000 ml , the sample can be pressed through a narrow valve under high pressure ( Manton-Gaulin homogenizer ). Cavitation forces also act here. The digestion under pressure on a small scale is done using a French press , which works on the same principle as the Manton-Gaulin homogenizer.
- A process principle similar to high pressure homogenization is used in the microfluidizer technology. In the Microfluidizer, the cell suspension is guided under high pressure through a microchannel with a fixed internal geometry, in which the cell walls are broken up due to shear forces and impact effects. The Microfluidizer can be used from sample quantities of 1 ml up to several 100 l / h.
- With the nitrogen decompression method, animal and plant cells as well as sensitive bacteria can be broken down quickly and gently, regardless of the sample size. According to Henry's Law, nitrogen is enriched in the cells at higher gas pressures . A sudden pressure release leads to the bursting of the cell membranes .
- Repeated freezing and thawing ( freeze- thaw ) destroy the cells through the shear forces of the ice crystals and perforation. Treatment with hypotonic buffer solutions results in hypotonic lysis of the cells. However, the combination of these methods does not lead to complete homogenization.
Non-mechanical digestion process
Non-mechanical disruption methods are used for cells that cannot be easily broken (for example yeast ). They are usually gentler than the mechanical processes.
- In the case of non-plant cells, variants of phenol-chloroform extraction such as Trizol can be used to dissolve the membrane lipids from the cell membrane , but in the presence of guanidinium thiocyanate it is denaturing .
- Autolysis with toluene is induced in yeast cells , which creates holes in the cell membrane . The enzymatic lysis with Zymolyase destroys the glucan - cell wall , while Triton X-100 destroys the cell membrane.
- In the case of Gram-positive bacteria, treatment with lysozyme destroys the peptidoglycan envelope , then the cell membrane can be loosened with Triton X-100 .
- In Gram-negative bacteria, the lipopolysaccharide of the outer cell membrane is dissolved by treatment with EDTA , then the peptidoglycan envelope is destroyed by treatment with lysozyme, then the cell membrane can be destroyed with Triton X-100.
- In the case of Gram-negative bacteria, the membrane lipids can be saponified using alkaline lysis ; due to the high pH value , this method is also denaturing.
Cell fractionation
In eukaryotic cells, the different cell components are often fractionated by centrifugation .
Protein purification
When the cells are disrupted, the proteins enter an unphysiological environment and must be protected from inactivation, denaturation and proteolysis during protein purification . Since proteases are released during the digestion, fast work at low temperatures (close to 0 ° C) is necessary for higher yields.
Proteolysis
Protease inhibitors are added to avoid degradation of the proteins .
pH change
Due to the continued partial metabolism , the pH value can change. It is therefore important to ensure adequate buffering . It may be necessary to readjust the pH value by adding ammonia or tris (hydroxymethyl) aminomethane solution (Tris).
Ionic strength
An ionic strength of less than 0.05 M can lead to the formation of inclusion bodies and a low yield as proteins attach to each other and to cell debris. Therefore, the digestion buffer solution should contain 0.05–0.1 M sodium chloride or potassium chloride .
Hydrophobic effects
An aggregation of the proteins by Van-der-Waals interactions and the formation of inclusion bodies due to hydrophobic effects can be obtained by adding non -ionic detergents such as Triton X-100 (about 0.02% w / v) or Lubrol PX (ca. 0.006% w / v).
Thiol oxidation
The oxidation of thiol groups can be avoided by adding dithiothreitol (DTT), dithioerythritol (DTE), 2-mercaptoethanol or tris (2-carboxyethyl) phosphine to the buffer solution (in single-digit mM concentrations). This is particularly necessary with cytosolic proteins, since a reducing environment prevails there.
Heavy metal ions
Heavy metal ions can react with functional groups on the protein. Bivalent cations ( Ca 2+ , Mg 2+ ) activate metalloproteases , therefore ethylenediaminetetraacetic acid (EDTA) is added in mM concentrations, which forms complexes with metal ions.
DNA purification
After cell disruption, various methods can be used to purify the DNA. To inactivate deoxyribonucleases , EDTA is used as a chelator of their cofactor ( magnesium ions ).
RNA purification
The methods of RNA extraction are similar to those of DNA extraction due to the similarity of the molecules.
literature
- Friedrich Lottspeich , Haralabos Zorbas: Bioanalytics . Spektrum Akademischer Verlag, Heidelberg 1998, ISBN 3-8274-0041-4 .
- Hubert Rehm , Thomas Letzel: The Experimenter: Protein Biochemistry / Proteomics. 6th edition. Spectrum Akademischer Verlag, Heidelberg 2009, ISBN 978-3-8274-2312-2 .
Web links
Individual evidence
- ↑ D. Freifelder, PF Davison: Studies on the sonic degradation of deoxyribonucleic acid. In: Biophys J. (1962), Vol. 2, pp. 235-247. PMID 13894963 ; PMC 1366369 (free full text).
- ↑ P. Chomczynski, N. Sacchi: Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. In: Anal Biochem . (1987) 162 (1), pp. 156-159. PMID 2440339 .
- ↑ HC Birnboim, J. Doly: A rapid alkaline extraction procedure for screening recombinant plasmid DNA. In: Nucleic Acids Res. (1979), Vol. 7 (6), pp. 1513-1523. PMID 388356 ; PMC 342324 (free full text).