Nodaviridae: Difference between revisions
No edit summary |
Added s2cid. Removed proxy/dead URL that duplicated identifier. |
||
| (46 intermediate revisions by 25 users not shown) | |||
| Line 1: | Line 1: | ||
{{Short description|Family of viruses}} |
|||
{{taxobox |
|||
{{Use dmy dates|date=April 2017}} |
|||
| virus_group = iv |
|||
{{Virusbox |
|||
| ⚫ | |||
| image = Journal.ppat.1005203.g001.D.jpg |
|||
| image_alt = |
|||
| image_caption = [[Crystal structure]] of grouper nervous necrosis virus a [[fish|piscine]] [[betanodavirus]] |
|||
| ⚫ | |||
| subdivision_ranks = Genera |
| subdivision_ranks = Genera |
||
| subdivision = |
| subdivision = |
||
| ⚫ | |||
| ⚫ | |||
}} |
}} |
||
'''''Nodaviridae''''' is a family of [[ |
'''''Nodaviridae''''' is a family of [[nonenveloped]] [[positive-strand RNA virus]]es.<ref>{{cite journal |last1=Sahul Hameed |first1=AS |last2=Ninawe |first2=AS |last3=Nakai |first3=T |last4=Chi |first4=SC |last5=Johnson |first5=KL |last6=ICTV Report |first6=Consortium |title=ICTV Virus Taxonomy Profile: Nodaviridae. |journal=The Journal of General Virology |date=January 2019 |volume=100 |issue=1 |pages=3–4 |doi=10.1099/jgv.0.001170 |pmid=30431412|doi-access=free |s2cid=53435394 }}</ref> Vertebrates and invertebrates serve as natural hosts. Diseases associated with this family include: viral [[encephalopathy]] and [[retinopathy]] in fish. There are nine species in the family, assigned to two genera.<ref name= ICTV>{{cite web |title=ICTV Report Nodaviridae |url=http://www.ictv.global/report/nodaviridae}}</ref><ref name=ViralZone>{{cite web|title=Viral Zone|url=http://viralzone.expasy.org/all_by_species/47.html|publisher=ExPASy|access-date=15 June 2015}}</ref> |
||
== |
==History== |
||
The name of the family is derived from the [[Japan]]ese village of Nodamura, [[Iwate Prefecture]] where Nodamura virus was first isolated from ''[[Culex tritaeniorhynchus]]'' mosquitoes.{{cn|date=October 2022}} |
|||
== Virology == |
|||
| ⚫ | |||
===Structure=== |
|||
| ⚫ | |||
| ⚫ | |||
=== Genome === |
|||
| ⚫ | RNA1, which is ~3.1 kilobases in length, |
||
[[File:Elife-25940-fig1-v2.jpg|thumb|[[Flock House virus]] genome and functional map of [[RNA-dependent RNA polymerase|replicase]] protein A.]] |
|||
| ⚫ | |||
| ⚫ | RNA1, which is ~3.1 kilobases in length, encodes a protein that has multiple functional domains: a mitochondrial targeting domain, a transmembrane domain, an RNA-dependent RNA polymerase (RdRp) domain, a self-interaction domain and an RNA capping domain. In addition, RNA1 encodes a subgenomic RNA3 that encodes protein B2, an RNA silencing inhibitor.<ref name=ICTV/> |
||
| ⚫ | |||
{| class="wikitable sortable" style="text-align:center" |
|||
|- |
|||
! Genus !! Structure || Symmetry !! Capsid !! Genomic Arrangement !! Genomic Segmentation |
|||
|- |
|||
|Betanodavirus||Icosahedral||T=3||Non-Enveloped||Linear||Segmented |
|||
|- |
|||
|Alphanodavirus||Icosahedral||T=3||Non-Enveloped||Linear||Segmented |
|||
|} |
|||
| ⚫ | |||
| ⚫ | |||
| ⚫ | Viral replication is cytoplasmic. Entry into the host cell is achieved by penetration into the host cell. Replication follows the positive stranded RNA virus replication model. Positive stranded |
||
| ⚫ | |||
{| class="wikitable sortable" style="text-align:center" |
|||
| ⚫ | Viral replication is cytoplasmic. Entry into the host cell is achieved by penetration into the host cell. Replication follows the positive stranded RNA virus replication model. Positive stranded RNA virus transcription, using the internal initiation model of subgenomic RNA transcription is the method of transcription. Vertebrates and invertebrates serve as the natural host. Transmission routes are contact and contamination.<ref name=ICTV/><ref name=ViralZone /> |
||
|- |
|||
! Genus !! Host Details !! Tissue Tropism !! Entry Details !! Release Details !! Replication Site !! Assembly Site !! Transmission |
|||
|- |
|||
|Betanodavirus||Fish||None||Unknown||Lysis||Cytoplasm||Cytoplasm||Passive diffusion, direct contact |
|||
|- |
|||
|Alphanodavirus||Insects, mammals, fishes.||None||Unknown||Lysis||Cytoplasm||Cytoplasm||Unknown |
|||
|} |
|||
==Taxonomy== |
==Taxonomy== |
||
| ⚫ | The members of the genus ''Alphanodavirus'' were originally isolated from insects while those of the genus ''Betanodavirus'' were isolated from fish. A small number of nodoviruses seem to lie outside either of these clades.<ref name=ICTV/> ''[[Flock house virus]]'' (FHV) is the best studied of the nodaviruses.<ref name="ICTV" /> There are nine species in this family, assigned to two genera:<ref name="ICTV" /> |
||
{{Div col|colwidth=20em}} |
|||
| ⚫ | |||
| ⚫ | |||
** ''[[Black beetle virus]]'' |
|||
<big>'''Group: ssRNA(+)'''</big> |
|||
| ⚫ | |||
{{Collapsible list|title= <big>Order: Unassigned</big> |
|||
** ''[[Flock House virus]]'' |
|||
|1={{Collapsible list| framestyle=border:none; padding:1.0em;|title=Family: Nodaviridae |
|||
** ''[[Nodamura virus]]'' |
|||
|1={{hidden begin|title=<small>Genus: [[Alphanodavirus]]</small>}} |
|||
* |
** ''[[Pariacoto virus]]'' |
||
| ⚫ | |||
| ⚫ | |||
* |
** ''[[Barfin flounder nervous necrosis virus]]'' |
||
* |
** ''[[Redspotted grouper nervous necrosis virus]]'' |
||
| ⚫ | |||
*<small>[[Pariacoto virus]]</small> |
|||
| ⚫ | |||
{{ |
{{div col end}} |
||
|2={{hidden begin|title=<small>Genus: [[Betanodavirus]]</small>}} |
|||
*<small>[[Barfin flounder nervous necrosis virus]]</small> |
|||
*<small>[[Redspotted grouper nervous necrosis virus]]</small> |
|||
| ⚫ | |||
| ⚫ | |||
{{hidden end}} |
|||
}} |
|||
}}<ref name=ICTV /> |
|||
While NoV remains the type species for this group, ''[[Flock house virus]]'' (FHV) is the best studied of the Nodaviruses.<ref name=ICTVdb>[http://phene.cpmc.columbia.edu/ICTVdB/WWW/00045FAM.htm International Committee on Taxonomy of Viruses]</ref><ref name=Leicester>[http://www-micro.msb.le.ac.uk/3035/VirusGroups.html University of Leicester]</ref> |
|||
==References== |
==References== |
||
| Line 69: | Line 52: | ||
==External links== |
==External links== |
||
* [http://www.ictv.global/report/nodaviridae '''ICTV Report:''' ''Nodaviridae''] |
|||
* [http://viralzone.expasy.org/all_by_species/47.html '''Viralzone''': Nodaviridae] |
* [http://viralzone.expasy.org/all_by_species/47.html '''Viralzone''': Nodaviridae] |
||
* [http://ictvonline.org/virusTaxonomy.asp '''ICTV'''] |
|||
{{Baltimore classification}} |
{{Baltimore classification}} |
||
{{Taxonbar|from=Q3772908}} |
|||
| ⚫ | |||
[[Category:Nodaviridae]] |
[[Category:Nodaviridae| ]] |
||
[[Category:Virus families]] |
|||
[[Category:Fish viral diseases]] |
|||
[[Category:Insect viral diseases]] |
|||
| ⚫ | |||
Latest revision as of 13:00, 16 January 2024
| Nodaviridae | |
|---|---|

| |
| Crystal structure of grouper nervous necrosis virus a piscine betanodavirus | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Kitrinoviricota |
| Class: | Magsaviricetes |
| Order: | Nodamuvirales |
| Family: | Nodaviridae |
Nodaviridae is a family of nonenveloped positive-strand RNA viruses.[1] Vertebrates and invertebrates serve as natural hosts. Diseases associated with this family include: viral encephalopathy and retinopathy in fish. There are nine species in the family, assigned to two genera.[2][3]
History[edit]
The name of the family is derived from the Japanese village of Nodamura, Iwate Prefecture where Nodamura virus was first isolated from Culex tritaeniorhynchus mosquitoes.[citation needed]
Virology[edit]
Structure[edit]
The virus is not enveloped and has an icosahedral capsid (triangulation number = 3) ranging from 29 to 35 nm in diameter. The capsid is constructed of 32 capsomers.[2]
Genome[edit]
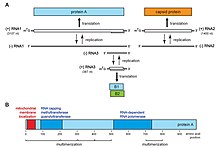
The genome is linear, positive sense, bipartite (composed of two segments – RNA1 and RNA2) single stranded RNA consisting of 4500 nucleotides with a 5’ terminal methylated cap and a non-polyadenylated 3’ terminal.[2]
RNA1, which is ~3.1 kilobases in length, encodes a protein that has multiple functional domains: a mitochondrial targeting domain, a transmembrane domain, an RNA-dependent RNA polymerase (RdRp) domain, a self-interaction domain and an RNA capping domain. In addition, RNA1 encodes a subgenomic RNA3 that encodes protein B2, an RNA silencing inhibitor.[2]
RNA2 encodes protein α, a viral capsid protein precursor, which is auto-cleaved into two mature proteins, a 38 kDa β protein and a 5 kDa γ protein, at a conserved Asn/Ala site during virus assembly.[2]
Life cycle[edit]
Viral replication is cytoplasmic. Entry into the host cell is achieved by penetration into the host cell. Replication follows the positive stranded RNA virus replication model. Positive stranded RNA virus transcription, using the internal initiation model of subgenomic RNA transcription is the method of transcription. Vertebrates and invertebrates serve as the natural host. Transmission routes are contact and contamination.[2][3]
Taxonomy[edit]
The members of the genus Alphanodavirus were originally isolated from insects while those of the genus Betanodavirus were isolated from fish. A small number of nodoviruses seem to lie outside either of these clades.[2] Flock house virus (FHV) is the best studied of the nodaviruses.[2] There are nine species in this family, assigned to two genera:[2]
References[edit]
- ^ Sahul Hameed, AS; Ninawe, AS; Nakai, T; Chi, SC; Johnson, KL; ICTV Report, Consortium (January 2019). "ICTV Virus Taxonomy Profile: Nodaviridae". The Journal of General Virology. 100 (1): 3–4. doi:10.1099/jgv.0.001170. PMID 30431412. S2CID 53435394.
- ^ a b c d e f g h i "ICTV Report Nodaviridae".
- ^ a b "Viral Zone". ExPASy. Retrieved 15 June 2015.
