Polymerase chain reaction
The polymerase chain reaction ( English polymerase chain reaction ( PCR ) ), to genetic material (a method DNA ) in vitro to reproduce. The enzyme DNA polymerase is used for this. The term chain reaction in this context means that the products of previous cycles serve as starting materials for the next cycle and thus enable exponential replication.
The PCR is performed in biological and medical laboratories, for example for the detection of hereditary diseases and viral infections , for creating and checking genetic fingerprints , for the cloning of genes and for paternity tests used. The method was developed by the biochemist Kary Mullis in 1983. In 1993 he was awarded the Nobel Prize in Chemistry for this. PCR is one of the most important methods of modern molecular biology today , and many scientific advances in this area (e.g. in the context of the human genome project ) would not have been possible without this method.
history

In the early 1970s, the Norwegian postdoc Kjell Kleppe came up with the idea of duplicating DNA using two flanking primers in the laboratory of Nobel Prize winner Har Gobind Khorana , but the idea was forgotten. The polymerase chain reaction itself was reinvented in 1983 by Kary Mullis . His intention was to develop a novel DNA synthesis method that artificially duplicates DNA by duplicating it repeatedly in several cycles using an enzyme called DNA polymerase . Seven years after he published his idea, Mullis was awarded the 1993 Nobel Prize in Chemistry .
DNA polymerase is found in all living things and, during replication, duplicates DNA before cells divide . To do this, it binds to a single strand of DNA and uses a short, complementary oligonucleotide ( primer ) to synthesize a strand that is complementary to it. The enzyme was used in vitro in Mullis' original PCR experiment . The double-stranded DNA was first separated into two single strands by heating to 96 ° C. At this temperature, the E. coli DNA polymerase I initially used was destroyed and therefore had to be added again after each heating. This sequence of work steps had to be repeated several dozen times in a row in order to achieve sufficient amplification . Mullis' original method was therefore very inefficient because it required a lot of time, large amounts of DNA polymerase, and constant attention.
The use of thermostable DNA polymerases , i.e. enzymes that retain their polymerase activity and do not denature even at temperatures of approximately 100 ° C. , brought about a decisive improvement in PCR technology . One of the first thermostable DNA polymerases was obtained from the thermophilic bacterium Thermus aquaticus , which lived in hot springs, and was named Taq polymerase . By using thermostable DNA polymerases, there was no longer any need to constantly add new polymerase, and the whole PCR process could be considerably simplified and automated.
Mullis was working for the Californian biotech company Cetus at the time and was settled with a premium of $ 10,000. Years later, Cetus sold the rights to the PCR method, including the patent for the Taq DNA polymerase he used, to Roche for $ 300 million. However, the enzyme Taq was already described in 1980 by the Russian researcher Kaledin. For this reason, after years of litigation by Roche, the patent for Taq has now been revoked. The US patents for the PCR technology itself expired in March 2005.
The Taq polymerase is still widely used. Its disadvantage is that it sometimes produces errors in copying the DNA, which leads to mutations in the DNA sequence. Polymerases such as Pwo and Pfu , which are derived from archaea , have a correction mechanism that significantly lowers the number of mutations in the copied DNA.
PCR in practice
PCR is used to replicate a short, precisely defined part of a DNA strand (if sufficient starting material is available, the RFLP method or its successor is also used to create a genetic fingerprint ). The to be replicated region of DNA is also called amplicon refers to the products produced during the PCR and amplified . This can be a gene or just part of a gene or non-coding DNA sequences. In contrast to living organisms, the PCR process can only copy relatively short sections of DNA. In a standard PCR, these can be DNA fragments up to about three thousand base pairs (3 kbp) in length. With the help of certain polymerase mixtures, other additives in the PCR and optimal conditions, fragments with a length of more than 20-40 kbp can be replicated, which is still much shorter than the chromosomal DNA of a eukaryotic cell. For example, a human cell contains around three billion base pairs per haploid genome .
In its current applications, PCR requires several basic components. These are:
- The original DNA containing the section to be replicated (template)
- Two primers to define the starting point of the DNA synthesis on the two single strands of the DNA, whereby the area to be replicated is limited on both sides.
- DNA polymerase that is not destroyed at high temperatures to replicate (copy) the specified section (e.g. Taq polymerase )
- Deoxyribonucleoside triphosphates, the building blocks for the DNA strand synthesized by DNA polymerase
- Mg 2+ ions, essential for the function of the polymerase, stabilize the attachment of the primers and form soluble complexes with the deoxyribonucleoside triphosphates
- Buffer solutions that ensure a suitable chemical environment for the DNA polymerase
The polymerase chain reaction takes place in a so-called thermal cycler . This machine heats and cools the reaction vessels in it precisely to the temperature required for the respective step. To prevent evaporation, a tightly sealing reaction vessel or a layer of oil is used on the reaction mixture. Any condensation in the lid of the vessel is prevented by a heatable device lid (over 100 ° C).
Real-time PCR or digital PCR is recommended if the PCR is to serve primarily as quantitative evidence .
The use of a pyrophosphatase can possibly increase the effectiveness of the PCR. The enzyme catalyzes the hydrolysis of the pyrophosphate split off from the nucleoside triphosphates to form orthophosphate . Pyrophosphate can act as an inhibitor in PCR.
example
A typical composition of a PCR reaction is given here as a general example. Many examples of the most diverse variations of PCR can be found in the scientific literature in various combinations:
1.0 µl DNA solution (100 ng / µl) 2.0 µl primer (usually two, each 1 µl) (10 pmol / µl) 4.0 µl 10 mmol deoxy nucleotides (dATP, dGTP, dCTP, dTTP), each dNTP 1 µl 5.0 µl 10X concentrated polymerase buffer solution 37.0 µl H 2 O 1.0 µl Pfu , Taq or other thermostable DNA polymerase (1–5 U / µl)
50.0 µl total volume
A 200 µl reaction vessel with the 50 µl mixture is placed in the thermal cycler. In addition to individual 200 µl reaction vessels, eight connected 200 µl reaction vessels or PCR plates for up to 96 batches of 200 µl or 384 batches of 50 µl each can be used as reaction vessels, depending on the sample insert or heating block of the thermal cycler . The plates are closed either with a rubber cover or a self-adhesive transparent film. The first-generation thermal cyclers did not have a heated lid, which is why the reaction batches of these devices were covered with mineral oil to avoid evaporation of water during the PCR .
Theoretical process
The PCR process consists of about 20–50 cycles, which are carried out in a thermal cycler . The following information is intended as a guide. Usually a PCR has to be optimized for the specific reaction. Each cycle consists of three steps (see figure below):
- Denaturation (melting): First, the double-stranded DNA is heated to 94–96 ° C in order to separate the strands. The hydrogen bonds that hold the two strands of DNA together are broken. In the first cycle, the DNA is often heated for a longer period of time (initialization) to ensure that both the starting DNA and the primers have completely separated from each other and that only single strands are present. Some (so-called hot start) polymerases have to be activated by an even longer initial heating phase (up to 15 minutes). It is then quickly cooled to 65 ° C to prevent the double helix from re-forming.
- Primer hybridization ( primer annealing ): The temperature is held for about 30 seconds at a value which allows specific annealing of primers to the DNA. The exact temperature is determined by the length and the sequence of the primers (or the appropriate nucleotides in the primer, if mutations are to be introduced by these = site-directed mutagenesis). If the temperature chosen is too low, the primers can under certain circumstances attach to sequences that are not one hundred percent complementary and thus lead to unspecific products (“ghost bands”). If the temperature is too high, the thermal movement of the primers may be so great that they cannot adhere properly, so that there is no or only inefficient product formation. The temperature, which largely excludes the two above-mentioned effects, is usually 5-10 ° C below the melting point of the primer sequences; this usually corresponds to a temperature of 55 to 65 ° C.
- Elongation (extending, polymerization, lengthening, amplification): Finally, the DNA polymerase fills in the missing strands with free nucleotides. It starts at the 3 'end of the attached primer and then follows the DNA strand. The primer is not removed again, it forms the beginning of the new single strand. The temperature depends on the working optimum of the DNA polymerase used (68–72 ° C). However, only very specific enzymes can work at these temperatures. The Taq polymerase is often used here. This step takes about 30 seconds for every 500 base pairs, but varies depending on the DNA polymerase used. The PCR products are then stable for many days, even at room temperature, so that further processing does not have to take place immediately. In many laboratories it has become established to cool the samples in the thermal cycler down to 4–8 ° C after the end of the PCR. However, many manufacturers advise against this, as condensation in the metal blockgreatly reducesthe service life of a cycler with a Peltier element .
- (A) The starting DNA is initially present as a double strand (arrow direction: 5 '→ 3').
- (B, C) After denaturation, the single strands are separated and the primers can bind.
- (C, D) The polymerase produces the opposite strand by extending the primers 5 '→ 3' (in the direction of the arrow). The products are still too long at one end. This can be explained by the fact that only a starting point (primer), but not an end point, is precisely defined.
- (E, F) In the next cycle, PCR products of the correct length are created for the first time - but the opposing strands are still too long.
- (G, H) In the third cycle, the PCR product is created for the first time as a double strand of the correct length (the other products are not shown in H).
- (H, I, J) In the following cycles, the desired products multiply exponentially (since they themselves serve as a template for further strand syntheses), while the undesired long products (see products of the first cycle) only increase linearly (only the DNA used serves as Matrix).
This is the theoretical ideal case, but in practice there are also a small number of fragments shorter than the desired target DNA. These short fragments accumulate especially in the late cycles and can also lead to incorrect PCR products due to a mismatch of the primers. For this reason, PCR reactions usually only run through about 30 cycles so that mainly DNA of the desired length and sequence is produced.
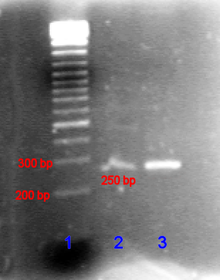
A PCR product can be identified by its size by agarose gel electrophoresis . (Agarose gel electrophoresis is a procedure in which DNA is placed in an agarose gel and a voltage is then applied. The shorter strands of DNA then move faster than the longer strands towards the positive pole.) The length of the PCR product can be determined by comparison with a DNA ladder that contains DNA fragments of known size and runs parallel to the sample in the gel.
PCR optimization
Various methods can be used to increase synthesis levels or to remove inhibitors of PCR.
PCR techniques
The classic PCR has been expanded and improved by numerous variations. This allows different tasks to be specifically approached. As an alternative to PCR, various methods of isothermal DNA amplification can also be used.
Real-time PCR (real-time PCR) or quantitative PCR (qPCR)
In laboratory jargon, almost only the English term real-time PCR or quantitative PCR is used, qPCR for short or rt-PCR for short, but this leads to confusion with the long-established term RT-PCR with upstream reverse transcription (see next section).
In real-time PCR, an initially inactive fluorescent dye is added to the reaction, which becomes active through the DNA production. (For example, because it is embedded in the DNA (like SYBR Green ) or because a quencher that initially quenches the fluorescence is removed during the amplification.) With each cycle - that is, "in real time" - the fluorescence is measured, which is the result one can infer the amount of amplified DNA.
Depending on the original number of copies, a certain threshold value of the fluorescence signal is reached sooner or later (or not at all). Because of this additional information, real-time PCR is nicknamed "quantitative PCR (qPCR)". For example, by using standard curves, this technique also allows absolute quantification (as the number of copies per reaction).
Although devices and dye-labeled reagents are more expensive than with “classic” endpoint PCR, it is not absolutely necessary to make the amplificates visible on a gel, so that work and, above all, time can be saved.
The qPCR technique can also be combined with an upstream reverse transcription (RT) , e.g. B. to detect RNA viruses , one speaks of qRT-PCR.
Reverse transcription and PCR (RT-PCR)
The amplification of RNA (e.g. a transcriptome or an RNA virus ) takes place in a reverse transcription PCR ( reverse transcription PCR ) via a reverse transcription of the RNA in DNA with a reverse transcriptase . In combination with a concentration determination in real time (qPCR), the reaction is called qRT-PCR.
Multiplex PCR
In multiplex PCR , more than one primer pair is used to amplify a particular gene or several genes at once. The multiplex PCR plays u. a. play a role in the diagnosis of diseases, for example in Lesch-Nyhan syndrome . Lesch-Nyhan syndrome is caused by a mutation in the HPRT1 gene, which codes for a hypoxanthine-guanine phosphoribosyltransferase . The gene has nine exons ; one of these exons may be missing in patients with Lesch-Nyhan syndrome. This can easily be discovered by a multiplex PCR.
Immunoquantitative real-time PCR (irt-PCR)
Sometimes even small amounts of pathogens have to be detected effectively, as they can also be dangerous for humans individually. The detection threshold of many immunological methods (e.g. ELISA ) can be insufficient for these cases, so that one falls back on immunoquantitative real-time PCR ( immunoquantitative real-time PCR ). This combines the high specificity of antibodies with a qPCR.
As with the classic ELISA, two antibodies are used . The first is fixed to a microtiter plate and recognizes the antigen being sought . The second antibody then binds to it. In conventional ELISA, the immunological complex of the first antibody, antigen and second antibody is made visible by a chemical color reaction, whereas in irt-PCR the second antibody is linked to a 246bp-long double-stranded DNA via a streptavidin - biotin complex. When the immunological complex is formed, this marker DNA can be amplified, detected and quantified by qPCR. This method is about a thousand times more sensitive than a classic ELISA.
Nested -PCR
The nested (nested) PCR is very suitable when only very small amounts of the DNA to be amplified are present relative to the total sample amount of DNA. Two PCRs are carried out one after the other. The first PCR - in addition to undesired sequence regions due to unspecific binding of the primers - generates the desired segment of the DNA (amplicon). The latter is used as a template for a second PCR. The desired sequence region is generated with very high specificity by means of primers that bind to areas within the first template ( downstream of the first primers). Since the DNA region of choice was also amplified for the second time, sufficient DNA is generated for further procedures.
The nested -PCR is used, for example, in genetic diagnostics, in forensics (for very little usable traces such as hair or blood drops, as in the criminal case JonBenét Ramsey ) or in phylogenetic examinations. Also microchimerism in leukocytes after a blood transfusion may be by means of nested be detected PCR.
Touchdown -PCR
The touchdown PCR avoids amplification of unspecific DNA sequences. In the first synthesis cycles, the annealing temperature is chosen to be just below the denaturation temperature. This means that the primer binding and thus also the amplificate are highly specific. The annealing temperature is reduced in the subsequent cycles. Although the primers can now enter into unspecific bindings, the specific replicates of the early reaction prevent excessive amplification of the unspecific sequences. Another advantage is an enormous increase in the amount of amplificate. This modified PCR thus allows a strong and very specific DNA amplification.
Digital PCR
In digital PCR (dPCR), the DNA is diluted and distributed over a large number of femtoliter reaction vessels. Either DNA is produced per reaction vessel or not. A digital signal is recorded . By counting a large number of reaction vessels, the proportion of reactions that have taken place can be used to determine the quantity.
application areas
PCR can be used for a variety of experiments and analyzes, some examples are presented below.
Genetic fingerprint
The genetic fingerprint is used in forensic genetics to identify a person by comparing their DNA with an existing sample. For example, a blood sample from a crime scene can be compared to the blood of a suspect. The sample can be minimal, i.e. a single cell (only white blood cells are possible, since there is no nucleus in red cells). At crime scenes, blood , semen , saliva , skin , hair or similar cells are usually found (although saliva and hair do not contain cell nuclei, cell remains are mostly found in saliva, and hair roots are often found in the hair). In theory, a single strand is sufficient. First you split the DNA sample into fragments, which are then amplified using PCR. The amplified fragments are then separated by gel electrophoresis. Since these fragments are multiplied by polymorphic areas ( polymorphisms ) that are highly variable, each person has a unique pattern, and this is known as the DNA fingerprint. Polymorphisms are in the non-coding area of DNA ( junk DNA ), so they do not reflect genes. Therefore, no disposition can be determined on the basis of the genetic fingerprint .
Paternity test

Although these generated “fingerprints” are unique ( they are almost identical in identical twins ; differences can only be detected with additional effort), genetic relationships, for example between parents and children or between siblings, can be determined by two or more genetic fingerprints what is used in a paternity test (see figure). A modification of this technique is also used to determine evolutionary relationships between organisms.
Disease detection
The detection of hereditary diseases in a given genome is a lengthy and complicated process, which can be shortened significantly by using PCR. Every gene that comes into question can be amplified (= duplicated) by PCR with the appropriate primers and then sequenced (sequencing DNA means determining the sequence of the nucleotides (or bases) of the DNA) in order to detect mutations .
Viral diseases can also be recognized by PCR by duplicating the virus DNA or, in the case of RNA viruses, first transcribing this RNA into DNA and then duplicating it using PCR ( RT-PCR ). This analysis can be done immediately after infection, often days or weeks before symptoms appear. If the diagnosis is made so early, it makes treatment much easier for the medical professionals. In addition, quantitative PCR is also used for diagnostics, e.g. B. to determine the exact viral load of a known HIV infection in order to understand the development of the therapy success.
The PCR can also be used for serial examinations. So she is z. B. used by blood donation services for routine blood tests. The so-called diagnostic window can be almost closed by the earliest possible discovery of a dangerous infectious disease in the donor (e.g. HIV , hepatitis B ) . The sensitivity of the PCR tests makes it possible to combine samples in so-called pools (e.g. 64 individual samples). If a pool tests positive, its size is reduced (usually halved) until the causative sample is found.
Cloning genes
Cloning a gene - not to be confused with cloning an entire organism - is a process in which a gene is isolated from one organism and then implanted in another. PCR is often used to amplify the gene, which is then inserted into a vector (a vector is a means by which a gene can be transplanted into an organism) such as a plasmid (a circular DNA molecule) (see Illustration). The DNA can then be used in another organism where the gene or its product can be better studied. The expressing a cloned gene can also for the mass production of useful proteins such. B. serve drugs.

(1) Chromosomal DNA from organism A.
(2) PCR.
(3) Multiple copies of a single gene from organism A.
(4) Insert the gene into a plasmid.
(5) Plasmid with the gene from organism A.
(6) Insertion of the plasmid into organism B.
(7) Duplication or expression of the gene from organism A in organism B.
Mutagenesis
Mutagenesis is a way of changing the sequence of nucleotides (bases) in DNA. There are situations in which you need mutated (changed) copies of a particular strand of DNA to determine the function of a gene. Mutations can be inserted into copied DNA sequences in two fundamentally different ways during the PCR process.
Targeted mutagenesis (for example "site-directed mutagenesis") allows the researcher to generate mutations at specific points on the DNA strand. Usually the desired mutation is integrated into the primers used for the PCR. In the case of targeted or site-specific mutagenesis , at least one of the primers is not one hundred percent identical to the DNA to which it attaches. Such a mutation is introduced into the DNA fragment during the amplification.
Random mutagenesis, on the other hand, is based on the use of error-prone polymerases (or polymerases without a mechanism for error correction) during the PCR process. With random mutagenesis, the location and type of mutations cannot be influenced and must first be identified by sequencing.
One application of random or targeted mutagenesis is the analysis of the structure-function relationships of a protein. After changing the DNA sequence, the resulting protein can be compared with the original and the function of all parts of the protein can be determined. In addition, functions of the nucleic acid itself (mRNA transport, mRNA localization, etc.) can be investigated.
Analysis of ancient (fossil) DNA
Since the PCR can generate any amount of material from only small amounts of DNA sample, it is particularly suitable for the very old aDNA , which in nature only occurs in amounts that are no longer sufficient for investigations. Almost all scientific knowledge gained in relation to aDNA and thus many long-extinct species are based on the method of PCR.
Sex determination
PCR can be used to obtain material for the detection of sex-specific chromosomes. This is not common with the classic domestic mammals, except possibly with hermaphrodites. For some animals in the zoo, but especially for birds, reptiles or fish, it is the gentlest and safest option. In the pet sector, this PCR method is common for parrots.
Food analysis
The increasing demands of the trade and official food control for the clarification and prevention of unfair competition led to the introduction of the technology into the food analysis . For example, PCR can be used to identify spices in complex food matrices. It can also be used to differentiate between varieties in fine cocoa z. B. Criollo and Forastero can be used.
variants
Various methods are based on the principle of PCR, e.g. B. DNA sequence analysis , ligase chain reaction , site-specific mutagenesis , ligation-during-amplification , megaprimer mutagenesis , inverse polymerase chain reaction , colony polymerase chain reaction , nested PCR , AFLP , MLPA , the MassTag-PCR , the immune polymerase chain reaction and the agglutination polymerase chain reaction .
literature
- For PCR (polymerase chain reaction)
- CR Newton, A. Graham: PCR. Introduction to Scientific Techniques . 2nd Edition. ed. BIOS Scientific Publishers, Oxford 1997, ISBN 1-872748-82-1 .
- RK Saiki, DH Gelfand, S. Stoffel, SJ Scharf, R. Higuchi, GT Horn, KB Mullis, HA Erlich: Primer-Directed Enzymatic Amplification of DNA with a Thermostable DNA Polymerase (PDF; 1.2 MB). In: Science . 239.1988, pp. 487-491, PMID 2448875 , ISSN 0036-8075
- Kary B. Mullis, F. Faloona, S. Scharf, R. Saiki, G. Horn, H. Erlich: Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction . Cold Spring Harb Symp Quant Biol. 1986; 51 Pt 1: 263-73.
- Kary B. Mullis, (et al.): System for automated performance of the polymerase chain reaction . United States Patent 5,656,493, August 12, 1997.
- Kary B. Mullis: The polymerase chain reaction. Birkhäuser, Boston 1994, ISBN 3-7643-3607-2 .
- Rabinow, Paul: Making PCR: A Story of Biotechnology , University of Chicago Press, 1996, ISBN 0-226-70146-8 .
- D. Baltimore: RNA-dependent DNA polymerase in virions of RNA tumor viruses . In: Nature 226, 1970, pp. 1209-1211.
- For genetic fingerprint ( genetic fingerprinting )
- J. Ballantyne (Ed.): DNA Technology and Forensic Science . Cold Spring Harbor Laboratory , Cold Spring Harbor NY 1989, ISBN 0-87969-232-4 .
- PR Billings (Ed.): DNA on Trial. Genetic Identification and Criminal Justice . Cold Spring Harbor Laboratory, Cold Spring Harbor NY 1992, ISBN 0-87969-379-7 .
- RK Saiki, S. Scharf, F. Faloona, KB Mullis, GT Horn, HA Erlich, N. Arnheim: Enzymatic Amplification of Beta-Globin Genomic Sequences and Restriction Site Analysis for Diagnosis of Sickle-Cell Anemia . (PDF; 1.5 MB) In: Science . 230.1985, pp. 1350-1354, ISSN 0036-8075 .
Web links
- Animation of the PCR with description (German)
- Flash animation of PCR (English)
- Website on practical aspects of real-time PCR and real-time RT-PCR (English)
- Tool for optimal calculation of annealing -Temperature of primers (English)
- Tool for creating appropriate primers (English)
- Film: "How does the PCR method work"
Individual evidence
- ^ K. Kleppe et al .: Studies on polynucleotides. XCVI. Repair replications of short synthetic DNAs as catalyzed by DNA polymerases. In: J. Mol. Biol. , 1971, Vol. 56, pp. 341-361, PMID 4927950 .
- ^ Information from the Nobel Foundation on the 1993 award ceremony to Kary B. Mullis (English) .
- ↑ Debunking the 4 degree myth: PCR can be left at room temperature overnight - miniPCR. Retrieved June 5, 2018 (American English).
- ^ A b Thomas M. Devlin: Textbook of Biochemistry with Clinical Correlations . John Wiley & Sons; 7th edition 2010, ISBN 978-0-470-28173-4 , p. 269.
- ↑ JS. Chamberlain et al .: Deletion screening of the Duchenne muscular dystrophy locus via multiplex DNA amplification . In: Nucleic Acids Res . 16 (23), 1988, pp. 11141–11156, PMID 3205741 , PMC 339001 (free full text)
- ↑ Bernard J. Glick, Jack J. Pasternak, Cheryl L. Patten: Molecular Biotechnology: Principles and Applications of Recombinant DNA . John Wiley & Sons, 4th edition 2010, ISBN 978-1-55581-498-4 , pp. 359ff.
- ↑ Cornel Mülhardt: The Experimenter: Molecular Biology / Genomics. 6th edition. Spektrum Akademischer Verlag, 2008, ISBN 978-3-8274-2036-7 , p. 99.
- ↑ Ulrich Busch: Molecular biological methods in food analysis: Basic methods and applications . Springer Verlag, Berlin 2010, ISBN 978-3-642-10715-3 , p. 113.
- ^ R Don, P Cox, B Wainwright, K Baker, J Mattick: Touchdown PCR to circumvent spurious priming during gene amplification . In: Nucleic Acids Res , 19 (14); 4008, PMID 1861999 , PMC 328507 (free full text)
- ↑ M. Fischer and I. Haase: PCR in der Lebensmittelanalytik, GIT Labor-Fachzeitschrift 3/06 ( 2006 ).
- ^ F. Focke, I. Haase and M. Fischer: Trends in food analysis: DNA-based ways to identify spices, Nachrichten aus der Chemie , 57, 1017-1020 (2009).
- ^ I. Haase, M. Fischer: Differentiation of Theobroma cacao and Theobroma grandiflorum by means of PCR . In: Journal for Consumer Protection and Food Safety . tape 2 , no. 4 , November 1, 2007, ISSN 1661-5751 , p. 422-428 , doi : 10.1007 / s00003-007-0252-1 .





