Nucleobases
| Purines | Pyrimidines | ||
 A denin |
|
||
 G uanine |
 C ytosin |
||
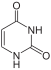
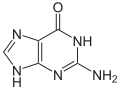
| Structural formulas of nucleobases in DNA ( A, G, C, T ) and RNA ( A, G, C, U ) - they are mostly bound via the NH group pointing down here. | |||
Nucleic bases , also known as nucleic bases , nucleobases or nucleobases , are part of nucleosides and nucleotides and thus the building blocks of nucleic acids in RNA and DNA .
They are called bases because they can be protonated on the nitrogen atoms and react weakly basic in aqueous solution . In the nucleic acids they are mostly N- glycosidically bound to ribose or deoxyribose . Base pairs can be formed via hydrogen bridges between nucleobases, which are structurally bearing in the double strand of DNA. The sequence of nucleobases in an RNA or DNA strand is also referred to as a base sequence .
The four bases adenine (A), guanine (G), cytosine (C) and thymine (T) occur in DNA , which is why they are also called DNA bases . In RNA, uracil (U) is used instead of thymine, accordingly A, G, C and U are also called RNA bases . Uracil differs from thymine only in the lack of a methyl group . The basic structure of uracil, thymine and cytosine is that of a pyrimidine , guanine and adenine are based on the basic structure of purine .
designation
The later Nobel Prize winner Albrecht Kossel (1853–1927) used the term nucleic bases as early as 1891 to describe the basic body, adenine, guanine and its derived from the decomposition of “nucleïn” and represented as cleavage products of the “nucleic acid” (from yeast) Derivatives, which I want to summarize all under the name of nuclein bases. ”In 1893 he discovered thymine, and in 1897 he also included cytosine under this term. In his Nobel lecture of 1910 on the chemical nature of the cell nucleus, he already understood the nucleic bases as the four nitrogen-rich "building blocks" which, together with two other different components - a carbohydrate and phosphoric acid - make up the nucleic acid molecule.
Occurrence
| base | Abbreviation | Occurrence |
|---|---|---|
| Adenine | A. | DNA, RNA |
| Guanine | G | DNA, RNA |
| Cytosine | C. | DNA, RNA |
| Thymine | T | DNA |
| Uracil | U | RNA |
| Hypoxanthine | HX | DNA, RNA |
| Xanthine | X | DNA, RNA |
The adjacent table lists the names, abbreviations and occurrences of nucleobases. As part of nucleosides and nucleotides , nucleobases have important functions. Together with ribose or deoxyribose , they form nucleosides, more precisely ribonucleosides or deoxyribonucleosides . With an additional phosphate group as nucleotides (ribonucleotides or deoxyribonucleotides), these form essential components of ribonucleic acid (RNA) and deoxyribonucleic acid (DNA), but are also contained in other important biomolecules.
Adenine, for example, occurs in adenosine in conjunction with a different number of phosphate groups as adenosine monophosphate (AMP), as cyclic adenosine monophosphate (cAMP), as adenosine diphosphate (ADP) and as adenosine triphosphate (ATP), in conjunction with nicotinamide in NADPH and NADH , and in connection with flavin in flavin adenine dinucleotide (FAD), as well as a part of coenzyme A . The same applies to guanine in guanosine triphosphate (GTP) and cytosine in cytidine triphosphate (CTP).
Hypoxanthine and xanthine are important intermediates in the synthesis of purines. They are neither regular components of DNA or RNA nor elements of the genetic code . However, under the action of mutagens - by deamination and the replacement of the amino group with a hydroxyl group and rearrangement into the tautomeric ketone - they can be formed from regular nucleobases: Hypoxanthine is formed from adenine , xanthine from guanine . In a similar way, uracil can also arise from cytosine .
structure
Purine bases
The basic structure of adenine, guanine, hypoxanthine and xanthine corresponds to the purine . This is why these molecules are also known as purine bases.
Pyrimidine bases
The basic structure of the bases cytosine, uracil and thymine is pyrimidine , which is why they are also referred to as pyrimidine bases.
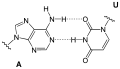
Base pairing

The purine base of one nucleotide can form a pair with a pyrimidine base of another nucleotide, which are linked by hydrogen bonds . Guanine (G) and cytosine (C) form such a base pair via three hydrogen bonds. Adenine (A) can form a base pair via two hydrogen bonds with thymine (T), as well as with uracil (U). The nucleic bases assigned to one another in these pairs are referred to as complementary bases.
The strand of a nucleic acid can also be connected to another nucleic acid strand by forming base pairs between its nucleotide building blocks. In this way, for example, two DNA strands can form a DNA double strand in which the complementary bases of one and the other strand are opposite each other (see double helix ). Similarly, base pairing can also be used to assign RNA nucleotides to the nucleotides of a single strand of DNA (see transcription ). Base pairs can also be formed between nucleotides of RNA strands, even intramolecularly in the same strand, so that strand sections are placed next to one another to form a hairpin structure .
With the four (DNA) bases occurring in DNA - G and A as well as C and T - the base pairs GC or CG and AT or TA can be formed complementarily paired.
With the four (RNA) bases G, A, C and U occurring in RNA, the base pair UA or AU is possible in complementary pairing in addition to GC and CG, rarely also as reverse pairing .
Building blocks of nucleic acids
In the nucleic acids, the nucleic occur depending bound to a sugar molecule with 5 C - atoms on a pentose , each on a phosphate group with two adjacent like pentoses esterified is. These via phosphodiester interconnected Pentosemoleküle form the backbone of a nucleic acid strand, which thus carries a number of different bases.
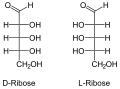
Nucleic acids are named after the type of their pentoses, which are present here in a ring as furanoses . Just like the ribose in RNA , the 2'- deoxyribose ( English deoxyribose ) does not occur naturally in DNA as an L - enantiomer . If the two β- depending anomers of D -Pentose installed, so β- D -Ribofuranose or β-2'-deoxy- D -ribofuranose . The latter does not have an OH group on the C2 'atom, so that it is only different by one missing oxygen atom .
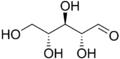
- Structural formulas of ribose and deoxyribose
D -Ribose in chain form
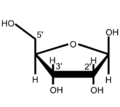
as a wedge line formulaβ- D- ribofuranose
as Haworth formula
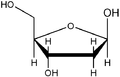
D - and L - deoxyribose
- Nucleosides
The compounds of nucleobase plus pentose are called nucleosides . With ribose as a monosaccharide and a name derived from the respective base, these include, for example, adenosine , guanosine , cytidine , thymidine and uridine . The nucleosides formed with deoxyribose are accordingly deoxyadenosine , deoxyguanosine , deoxycytidine , deoxythymidine and deoxyuridine . The nucleic base is linked to the pentose in a β-glycosidic bond . As a rule, this is done N-glycosidically on the nitrogen atom , i.e. in a 1-N-β-glycosidic bond. As an exception, there are also C-glycosidic bonds on the carbon atom of the nucleobase, such as the uracil in pseudouridine (Ψ), which can be found in the TΨC loop of a tRNA . Nucleosides are therefore glycosides , their aglycon is the nucleobase.
- Nucleotides

A phosphate group is bound to the pentose in the structural units of nucleic acids . These compounds of a nucleobase plus pentose plus phosphate are called nucleotides . The names of the nucleotides in RNA are derived from those of the nucleosides plus the final group of syllables - monophosphate . If the nucleotides are building blocks of DNA, then deoxy is prefixed, for example deoxyadenosine monophosphate , abbreviated dAMP.
Nucleic acids are polymeric macromolecules , more precisely polynucleotides . They are built up from reactive monomers by RNA polymerases and DNA polymerases . These nucleotides are nucleoside triphosphates, for example deoxyadenosine triphosphate , abbreviated dATP.
- Nucleic acids
RNA mostly occurs as a single polynucleotide strand, but can also form double strands by pairing complementary bases. More often, loop formations as a result of intramolecular pairing of strand sections which bear in opposite directions complementary sequences, also Palindrome mentioned.
In contrast, DNA usually does not consist of one polynucleotide strand, but of two, each of which is a chain of numerous nucleotides. The two strands are complementarily connected to one another via base pairs to form a double strand (see double helix ). An adenine is opposed to a thymine, a cytosine to a guanine.
The exact order of the four DNA bases of a strand is called the base sequence . Hereditary information is laid down and stored in the pattern of this base sequence . Certain sections code for the sequence of amino acids in the construction of proteins . DNA double strands can also be doubled or duplicated by adding another complementary strand to each strand, so that two identical double strands are created (see replication ).
- Base modifications
In addition to the listed primary nucleobases, various, rather rare, modifications occur naturally. In addition to the above -mentioned purine bases xanthine and hypoxanthine, these are modified bases such as 7-methylguanine or, as pyrimidine bases, 5-methylcytosine , 5-hydroxymethylcytosine and 5,6- dihydrouracil . With β- D -ribofuranose these bases form the corresponding nucleosides xanthosine , inosine , 7-methylguanosine , 5-methylcytidine , 5-hydroxymethylcytidine and dihydrouridine .
In addition, technical syntheses enable a large number of derivatives by introducing further substituents , as base analogues such as 5-fluorouracil or in xDNA or Hachimoji DNA .
See also
literature
- Jeremy M. Berg, John L. Tymoczko, Lubert Stryer : Biochemistry. 6 edition, Spektrum Akademischer Verlag, Heidelberg 2007. ISBN 978-3-8274-1800-5 .
- Donald Voet, Judith G. Voet: Biochemistry. 3rd edition, John Wiley & Sons, New York 2004. ISBN 0-471-19350-X .
- Bruce Alberts , Alexander Johnson, Peter Walter, Julian Lewis, Martin Raff, Keith Roberts: Molecular Biology of the Cell , 5th Edition, Taylor & Francis 2007, ISBN 978-0-8153-4106-2 .
Web links
Individual evidence
- ↑ Albrecht Kossel : About the chemical composition of the cell. (Lecture on January 30, 1891) In: Archives for Physiology. Year 1891, p. 184. online
- ^ Albrecht Kossel and Albert Neumann: About nucleic acid and thymic acid. In: Journal of Physiological Chemistry. Volume 22, No. 1, January 1897, p. 77. doi: 10.1515 / bchm2.1897.22.1.74
- ^ Albrecht Kossel: The Chemical Composition of the Cell Nucleus. Nobel Lecture, December 12, 1910.